Changelog
Version 3.15.4 (29-Apr-2026)
POSCAR / XDATCAR / CHGCAR file reader: Accept blank lines almost anywhere in a file.
pro Restored support for OpenSSH-based connection method to remote files via SFTP on Windows. The workaround is to use the Cygwin/MSYS2 build of sftp.exe that ships with Git for Windows instead of the incompatible version that comes with Windows.
Fixed the option to specify a filename pattern on the command line when launching OVITO, e.g.,
ovito 'data.*'. The single quotes ensure that the glob pattern is resolved by OVITO rather than by the shell.Fixed a bug in rendering of selected bonds and line elements with the VisRTX renderer in the interactive viewports.
Bug fix: Cancel buttons of Animation Key Editor dialog and Configure Trajectory Playback dialog do not restore original states.
Bug fix: Quitting the application via CMD+Q shortcut while a file selection dialog is open lets OVITO crash on macOS.
Version 3.15.3 (15-Apr-2026)
The new menu provides quick access to recently opened files and scenes.
The pipelines list widget in the upper right corner of the OVITO window now provides a button to toggle the visibility of individual pipelines in all viewports.
The redesigned modifier selector widget now supports keyboard input for quick item search: typing one or more characters highlights the first matching entry, and pressing Enter adds it. This restores a UI behavior that was present before the redesigned selector widget was introduced in OVITO 3.15.0.
The application settings now provide an option to switch from the operating system’s file selection dialog to the Qt widget-based file selection dialog, which can be useful on certain platforms (e.g. Linux with older GNOME desktops) where the native file dialog may ignore the working directory set by OVITO and always start in the same default folder.
Fixed a bug in rendering of bonds with fractional bond orders.
Fixed picking of bonds with bonds orders greater than 1 in the Manual selection modifier.
Updated dependencies: Python 3.13.13, Qt 6.10.3, pyside6 6.10.3, OpenSSL 3.0.20, libsodium 1.0.22, zlib 1.3.2, boost 1.90.0
Version 3.15.2 (31-Mar-2026)
Add support for double precision floats in the GSD/HOOMD file reader and GSD/HOOMD file exporter (issue).
Fixed list of pipelines sometimes not being updated after loading a .ovito state file (issue).
Version 3.15.1 (27-Mar-2026)
You can now specify an initial working directory for file imports when launching OVITO from the terminal.
The Surface mesh and Triangle mesh visual elements gained new options to control the width and color of wireframe lines rendered along mesh edges when the Highlight mesh edges option is enabled.
Gaussian Cube file reader: Added new option Convert density values from Bohr units to Angstroms, which controls whether the grid values are converted from \(\text{bohr}^{-3}\) units to \(\text{Å}^{-3}\) (OVITO’s internal units).
Fixed a bug that incorrectly labeled the central N atom of a nitro group in the Bond order modifier (reported here).
Fixed the display of bonds with bond order greater than 3 by clamping bond order values to the interval [0, 3].
Breaking change in the POSCAR / XDATCAR / CHGCAR file reader: The CHGCAR grid type has been changed from cell-data to point-data. This new interpretation is consistent with the grid definition in the VASP wiki.
Version 3.15.0 (03-Mar-2026)
OVITO 3.15 is a major release that brings powerful new modifiers for molecular modeling, a streamlined workflow for sharing and organizing modifiers, and broad improvements to file I/O, video rendering, and platform support.
Improved support for molecular structures
The Create bonds modifier gained a new Covalent Radius mode, which uses tabulated covalent radii to automatically determine bond cutoffs for each pair of chemical elements.
New modifier: Bond order pro – Automatically assigns bond orders (single, double, triple, aromatic) to molecular structures, enabling the visualization of double and triple bonds and the identification of functional groups. The Bonds visual element gained the ability to render different bond orders with distinct visual styles:
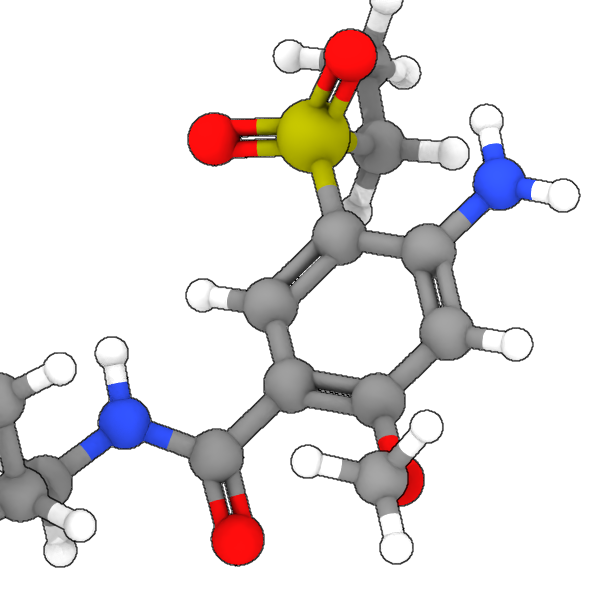
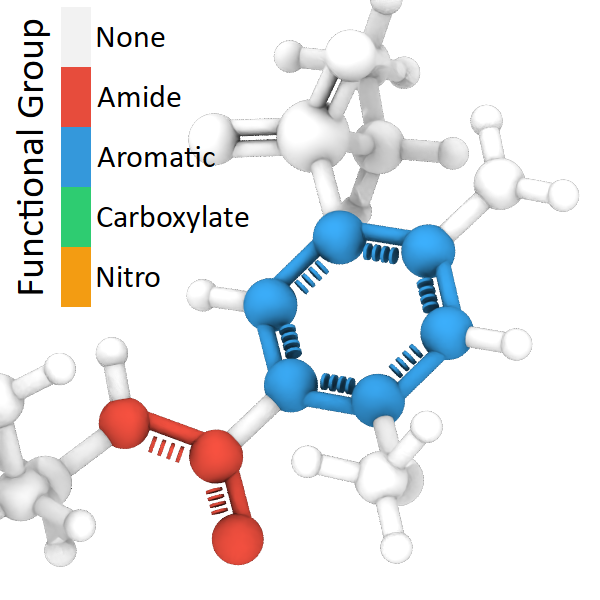
Modifier additions and changes
New modifier: Select overlapping particles – Detects and selects particles that overlap in space. Useful for cleaning up imported datasets or verifying simulation configurations.
The former Bond analysis modifier has been split into two focused modifiers, Bond angle distribution and Bond length distribution , for clearer functionality.
The former Coordination analysis modifier has been renamed to Radial distribution function (RDF) to better reflect its primary purpose.
The Expand selection modifier now offers a Molecule mode, which expands the current selection to include all particles belonging to the same molecule(s) – ideal for selecting entire molecules in complex systems.
Several modifiers now support processing only one particular object instead of the entire class of objects (e.g. surface meshes): Affine transformation, Delete selected, Replicate, and Slice.
The Assign color modifier now supports operating on Lines objects.
The value of the particle radius scaling factor is now remembered across program sessions.
Modifier snippets
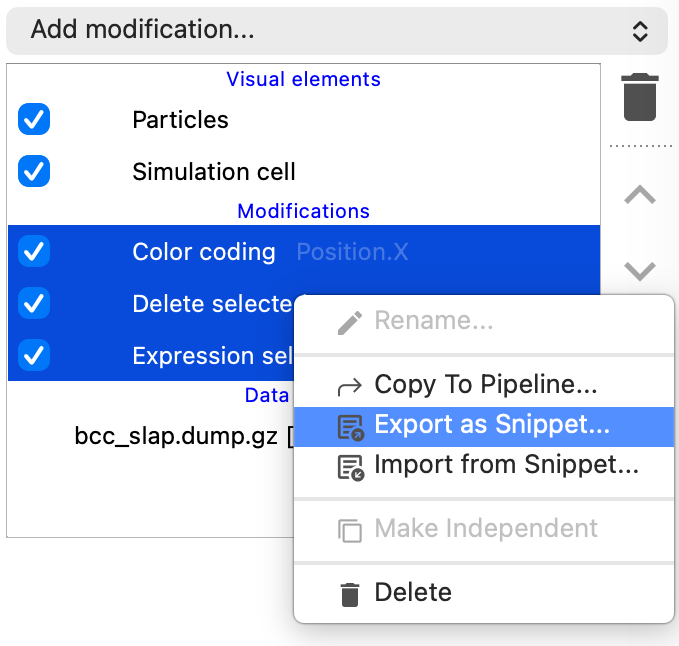
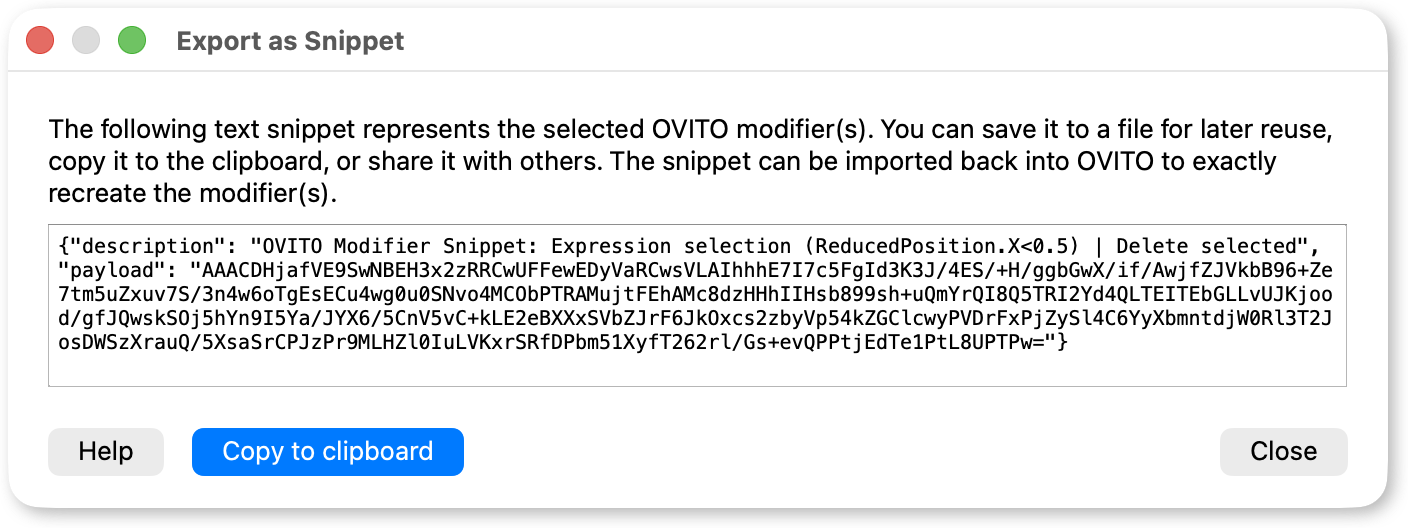
You can now export and import modifiers as text snippets via the context menu in the pipeline editor. This makes it easy to save your modifier sequences for later reuse, share them with colleagues, or transfer them between machines – all through a simple copy-and-paste of a text string.
New workflow for editing particle types and the simulation cell
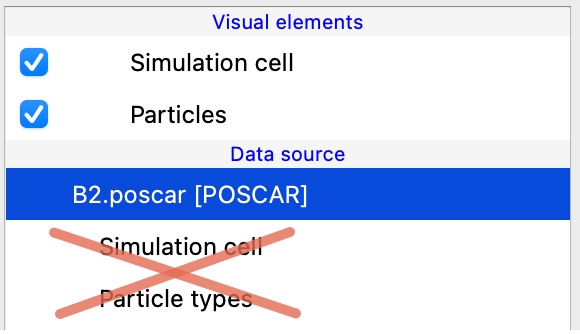
OVITO 3.15 removes the possibility to directly edit particle types and the simulation cell. The corresponding entries in the pipeline editor have been removed, because they bypassed the data pipeline system leading to unexpected behavior and confusion. Instead, two new modifiers will take over this job in the future, ensuring a clear, reproducible workflow.
New modifier: Edit simulation cell – Provides a dedicated, undo-able way to modify the simulation cell geometry and boundary conditions within the data pipeline. The new modifier also provides a more focused alternative to the Affine transformation modifier, which was previously often used for cell modifications.
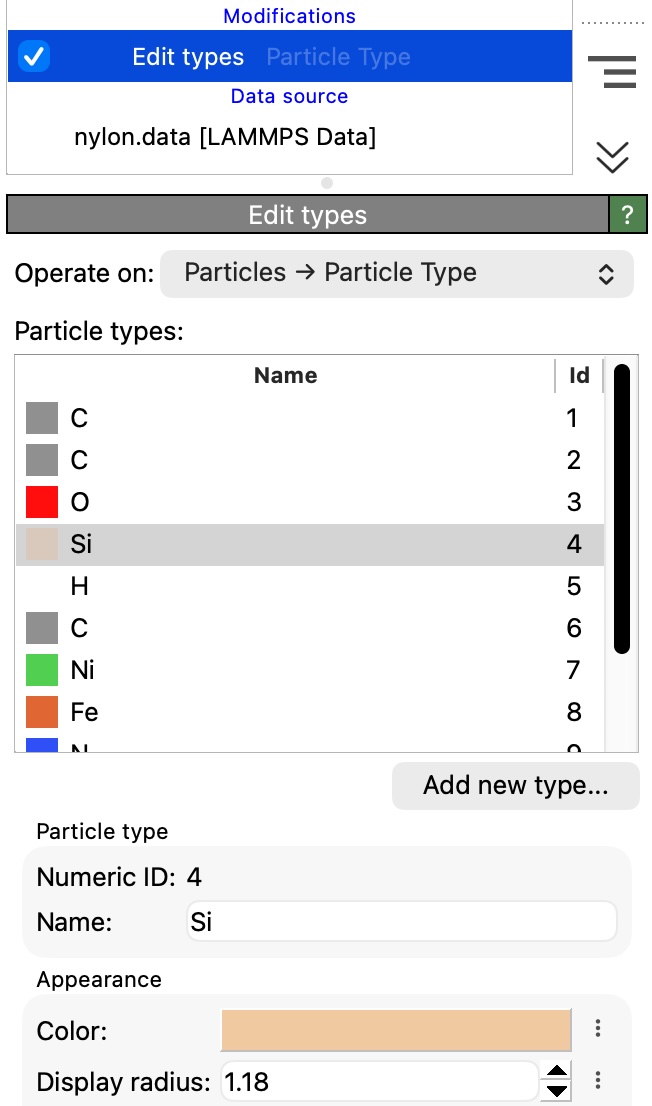
New modifier: Edit types – Lets you rename, recolor, and reshape particle, bond, or molecule types as a pipeline step, keeping type modifications visible and reproducible. For the first time, this modifier also provides a way to add and remove types, e.g., to introduce a new particle species in the dataset.
The new Types tab in the data inspector gives you a clear overview of all particle types, bond types, and other type classifications in your dataset:
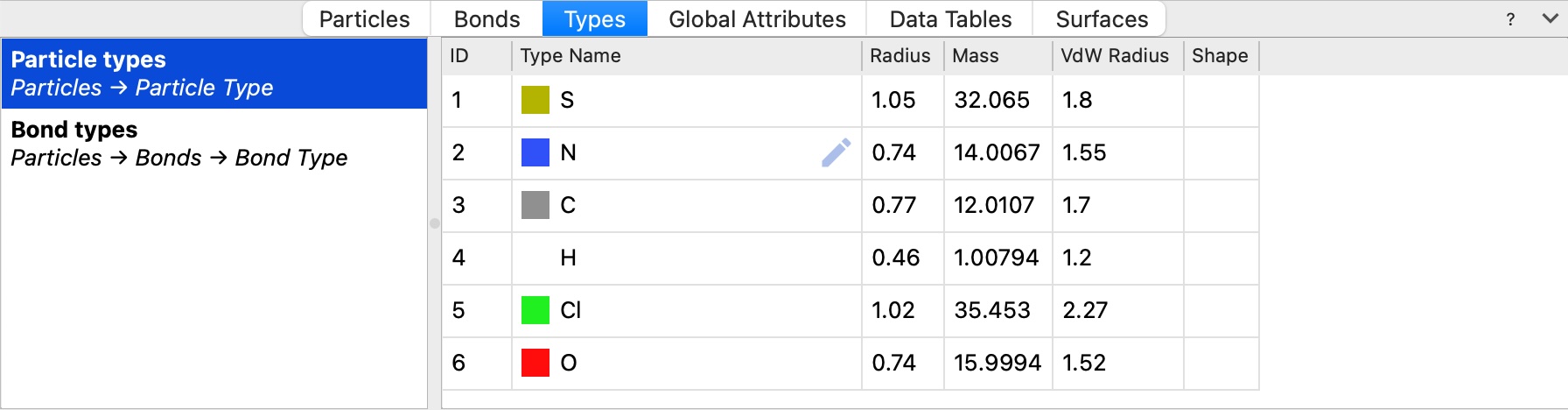
Click the pencil button, which appears when you hover over a type entry, to open the Edit types modifier with the corresponding type pre-selected for editing.
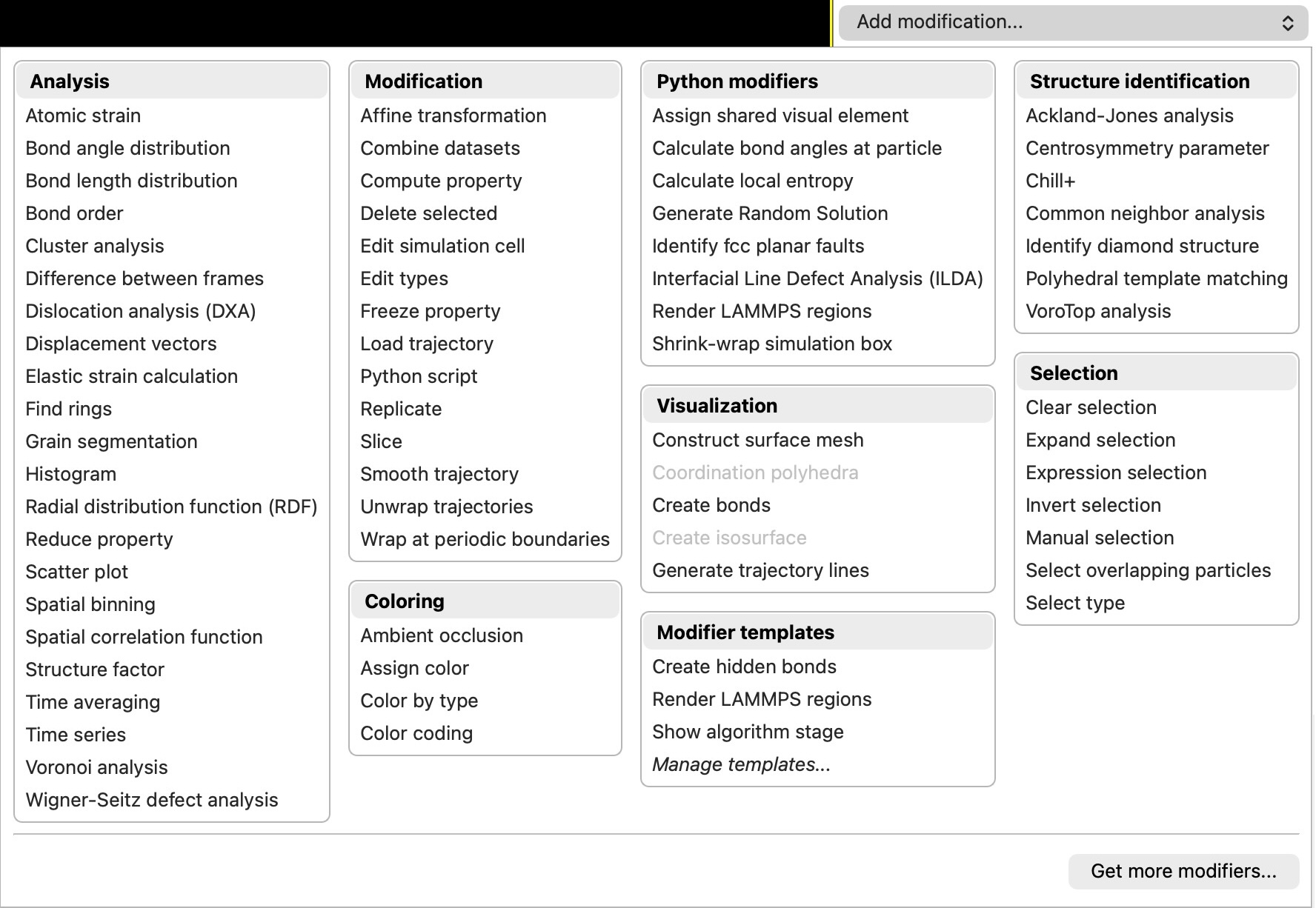
Redesigned modifier selector
The modifier selector widget has been redesigned with improved usability and a structured layout, making it easier to find and add modifiers to your pipeline:
New tutorial
Voronoi cell visualization – A step-by-step guide to visualizing Voronoi cells and exploring the local atomic neighborhood using OVITO
Video rendering
OVITO now supports external FFmpeg video encoding, giving you access to modern high-compression codecs such as H.264 and H.265 for smaller file sizes and better video quality:
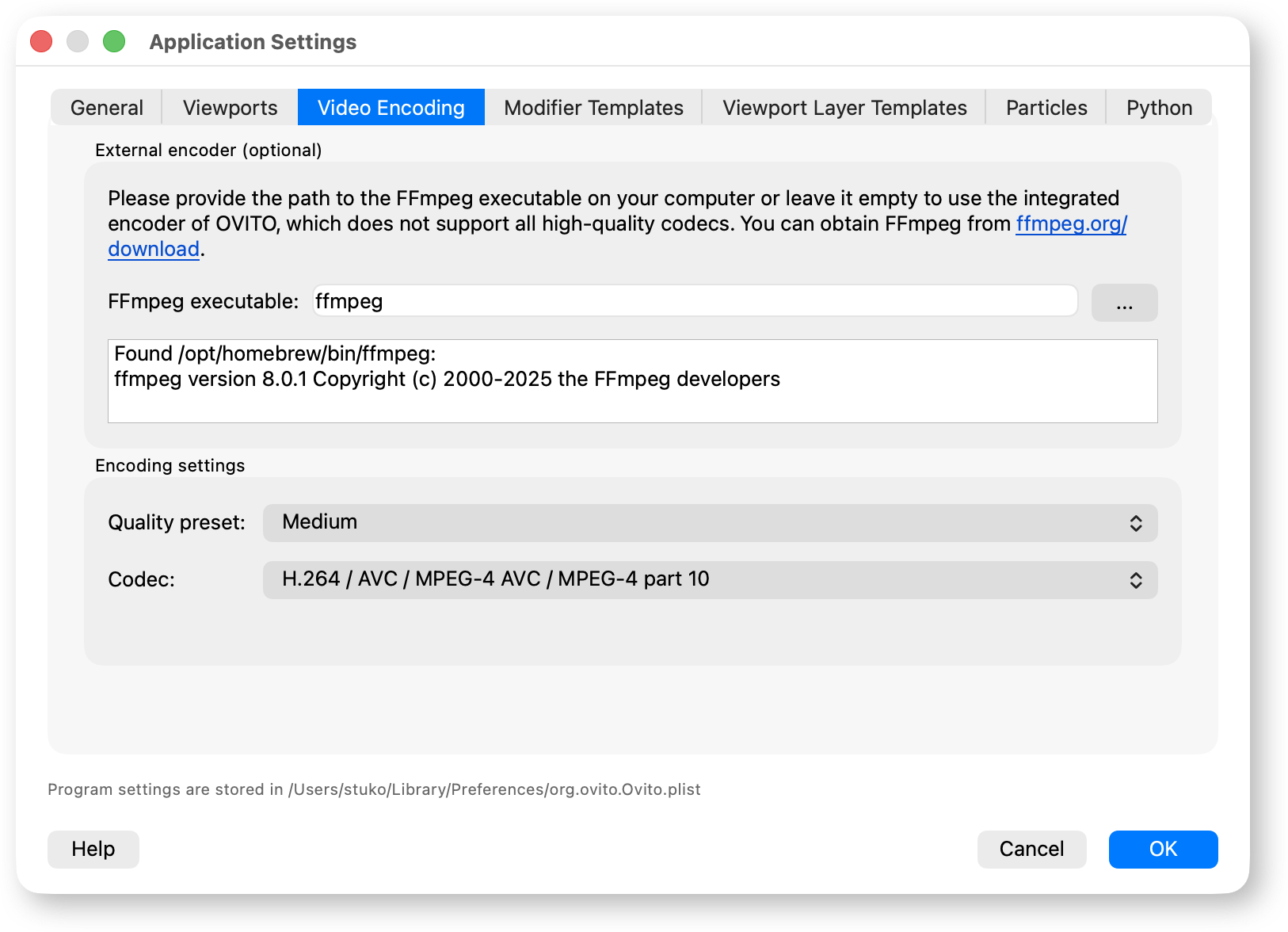
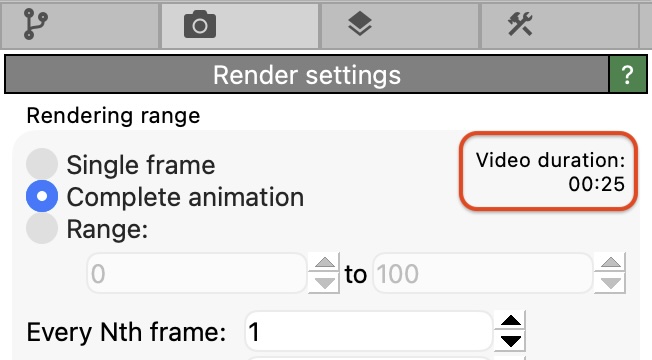
The render settings panel now displays a video duration estimate before rendering, which takes into account the number of trajectory frames and the configured playback frame rate:
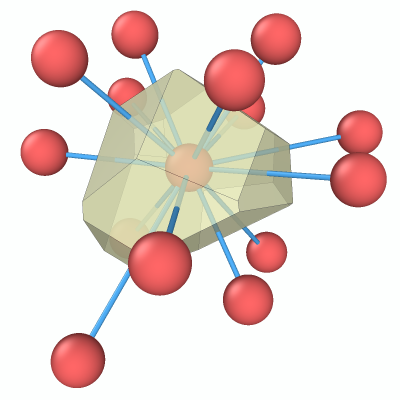
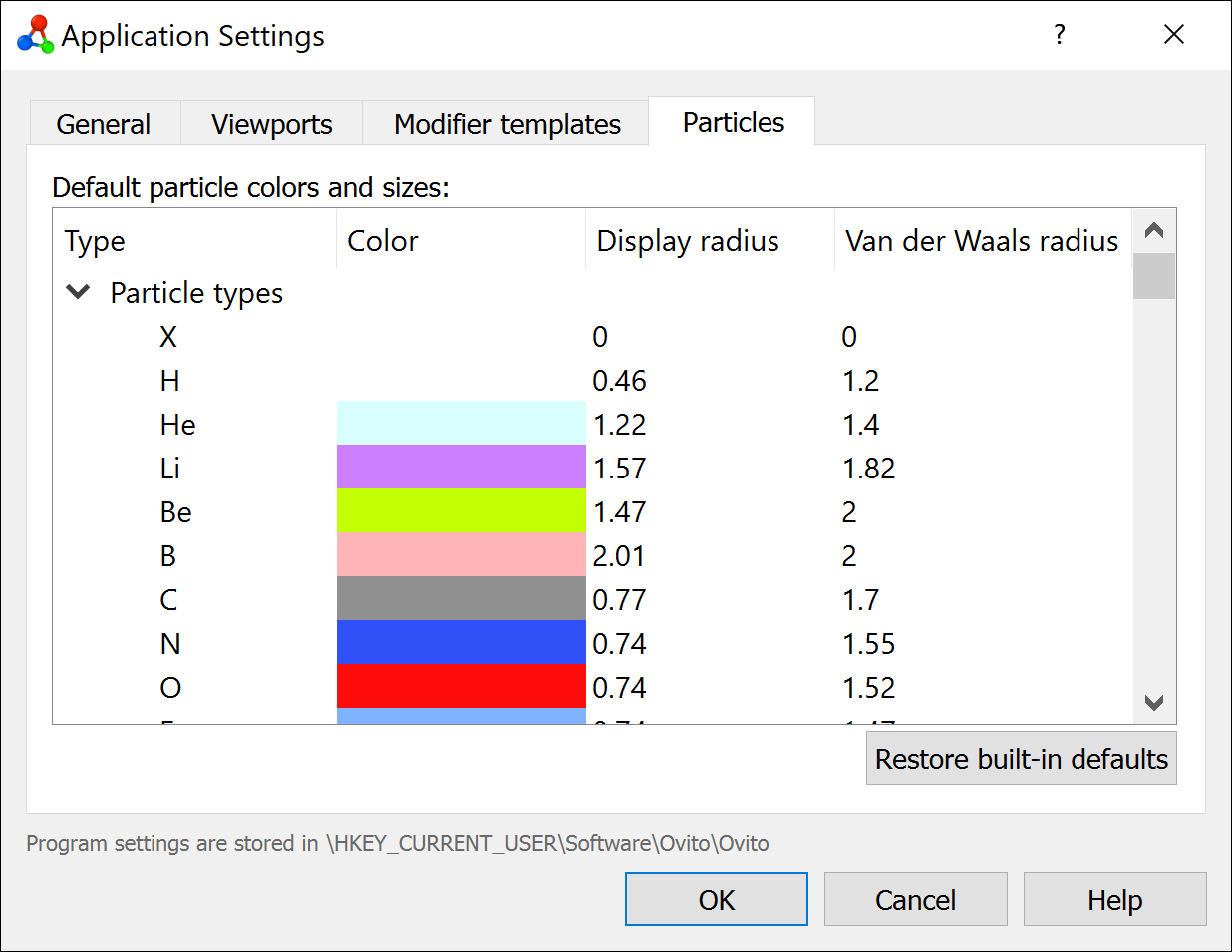
Chemical element themes
Customizable particle color and radius themes let you define your preferred visual defaults for chemical elements. Themes can now be exported and imported as JSON files, making it easy to share consistent visualization styles. We provide a theme for download that matches the color style of the VESTA atomistic visualization software.
File I/O improvements
Added a file reader for MOL/SDF files, commonly used for storing molecular structures in chemistry and drug discovery applications.
CIF file reader: Automatic creation of multi-element particle types for sites with partial occupancy at identical positions.
mmCIF file reader: Sequence IDs are now read as the particle property
Sequence.The PDB file reader now reads atom serial numbers as particle identifiers and
CONECTrecords as bonds.MercuryDPM file reader: Support for reading particle orientations stored as Euler angles, which get automatically converted to quaternions.
Aspherix file reader: Always create the
Particle typeproperty even if type information is missing in the file, allowing users to still assign custom particle shapes.Updated the gemmi library to version 0.7.3 for improved PDB/CIF/mmCIF file import.
Reduced the typical size of OVITO session state files for long trajectories.
Platform support
OVITO now runs natively on Linux aarch64 (ARM64 architecture).
OVITO installers now include a Software Bill of Materials (SBOM) documenting all third-party software components.
Interactive viewports
pro The VisRTX renderer now uses progressive refinement rendering in interactive viewports, delivering a more responsive experience when working with complex scenes.
Rendering issues in interactive viewports (e.g. exceeding the maximum number of particles supported by OpenGL) are now clearly indicated by an error icon.
Bug fixes
pro Fixed the Find rings modifier not finding rings of size equal to the configured maximum ring size.
pro Fixed the Find rings modifier reporting evenly sized rings in both directions.
pro Fixed broken “partition by bond/particle selection” modes in the former Bond analysis modifier (now replaced by the new Bond angle distribution and Bond length distribution modifiers).
pro VisRTX renderer: Fixed visual artifacts at semi-transparent object edges when compositing on light backgrounds.
py A warning is now issued when attempting to export an empty scene to a glTF file via
export_file().Fixed the PDB file reader’s Generate bounding box option not working.
Fixed detection of desktop dark color theme on Linux due to missing DBus module in Qt framework.
Version 3.14.1 (16-Oct-2025)
Add new simulation cell tab to OVITO’s data inspector.
Add filter expression fields to the Data Tables, Voxel Grids and Surfaces tabs of the data inspector, which allow filtering table rows based on a user-defined condition.
Data inspector tabs now display the total number of particles, bonds, etc. at the end of the pipeline.
Warn when user tries to render an animation without saving the result to disk.
Support more than 134M spherical particles in the OpenGL renderer (can now render up to 2.1B particles).
Fix: Filter expression variable
Frameis always 0 in data inspector.Fix: Tick labels in the timeline overlapping.
Add “°” suffix to angular parameter values.
Fix data inspector tab table width on windows.
Add ‘Help’ button to data inspector panel title bar.
Auto-disable the wildcard pattern option when user has imported a trajectory file to avoid accidentally concatenating multiple matching files into one long trajectory.
Fix: File source panel does not display the file-not-found error after opening a .ovito state file with missing input files.
Fix: File source title not updated in pipeline editor UI after picking a new simulation file.
Update dependencies: libssh 0.11.3, OpenSSL 3.0.18
pro List supported ANARI extensions in the system information dialog.
pro Disable the OpenSSH client on the Windows platform, because it is nonfunctional.
py New
PropertyContainer.append()method, which allows adding a new item to a property container and initializing its properties.py Accessing the attribute
PythonModifier.delegatenow auto-compiles the script code provided inPythonModifier.scriptand instantiate the customModifierInterfaceclass if necessary, e.g. after loading a pipeline from a .ovito state file.
Version 3.14.0 (21-Sep-2025)
Discrete color maps: A discretized color map can now be enabled to visualize integer properties with distinct color bands:
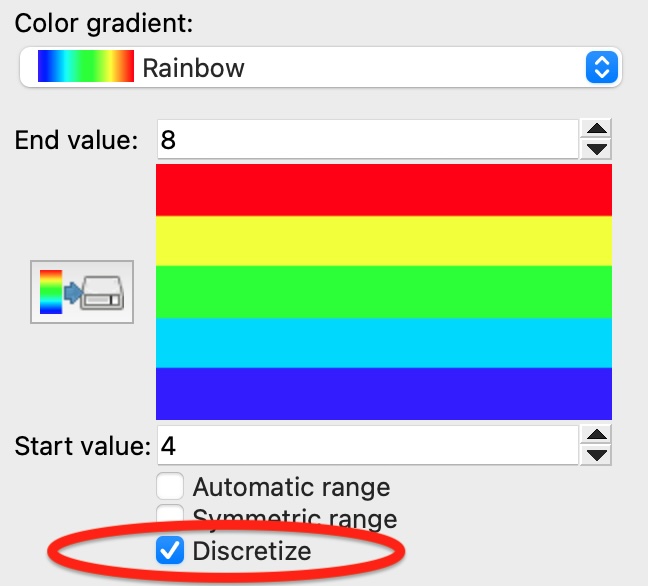
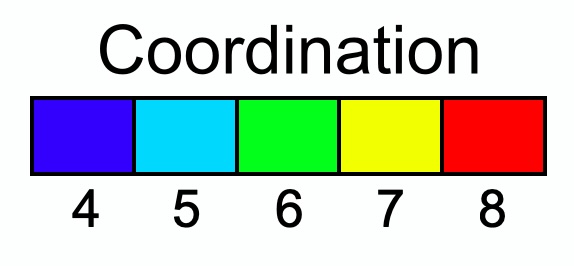
New modifier: Find rings pro - A high-performance algorithm to identify and visualize bond rings in molecular structures
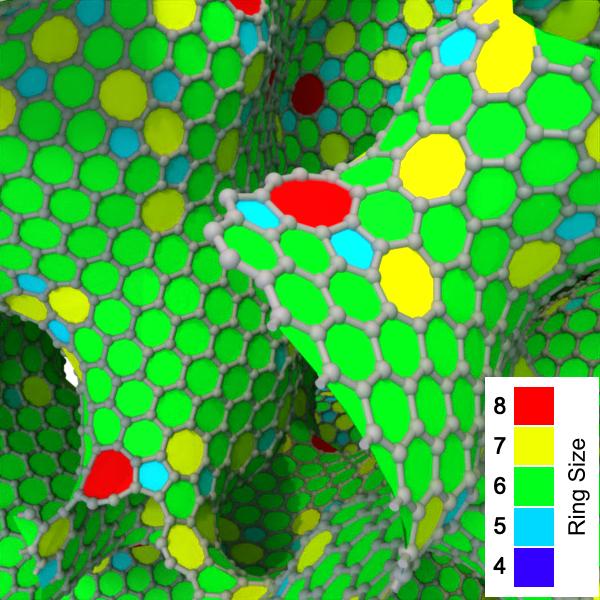
New modifier: Reduce property pro
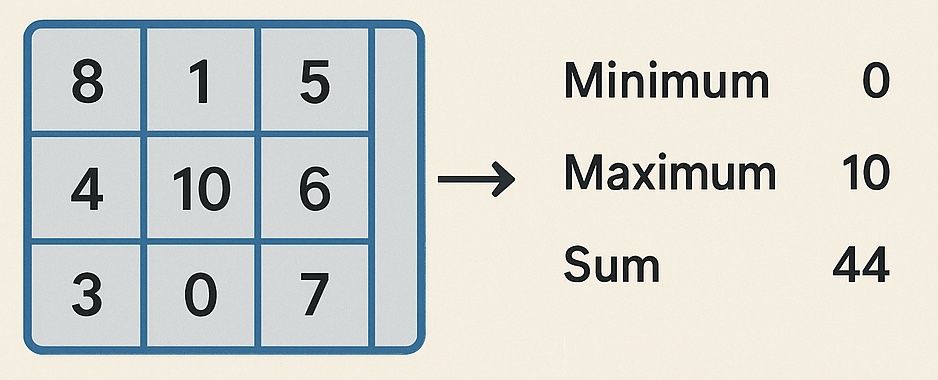
Reduces a selected data property to a single value by applying a specified reduction operation over all data elements. For example, the modifier allows computing the sum of a property across all particles.
The modifier offers the following reduction operations: minimum, maximum, sum, non-zero count, mean, median, variance, standard deviation, argmin, and argmax.
New modifier: Difference between frames pro - Computes the difference (delta) of a property between the current frame and a reference configuration. The reference configuration can be either fixed or relative to the current frame (time offset), enabling sliding-window difference calculations.
- The modifier can compute time differences for the following kinds of inputs:
Particle properties
Global attributes
Data tables
Voxel grid properties
New function available in the menu:
Send us your ideas and suggestions for new features or improvements in OVITO.
Radial distribution function (RDF) modifier:
The modifier can now count the number of neighbors of different types separately, which is useful for analyzing the chemical composition of the local neighborhood. Furthermore, it allows selecting other type classifications for the calculation of partial RDFs.
OVITO’s timeline can now display timesteps of the MD simulation instead of animation frame numbers:

Timestep display

Structure names for ASE databases
The new option can be enabled in the Animation settings dialog of OVITO.
The VisRTX renderer pro now lets you choose between the standard and physically-based material:
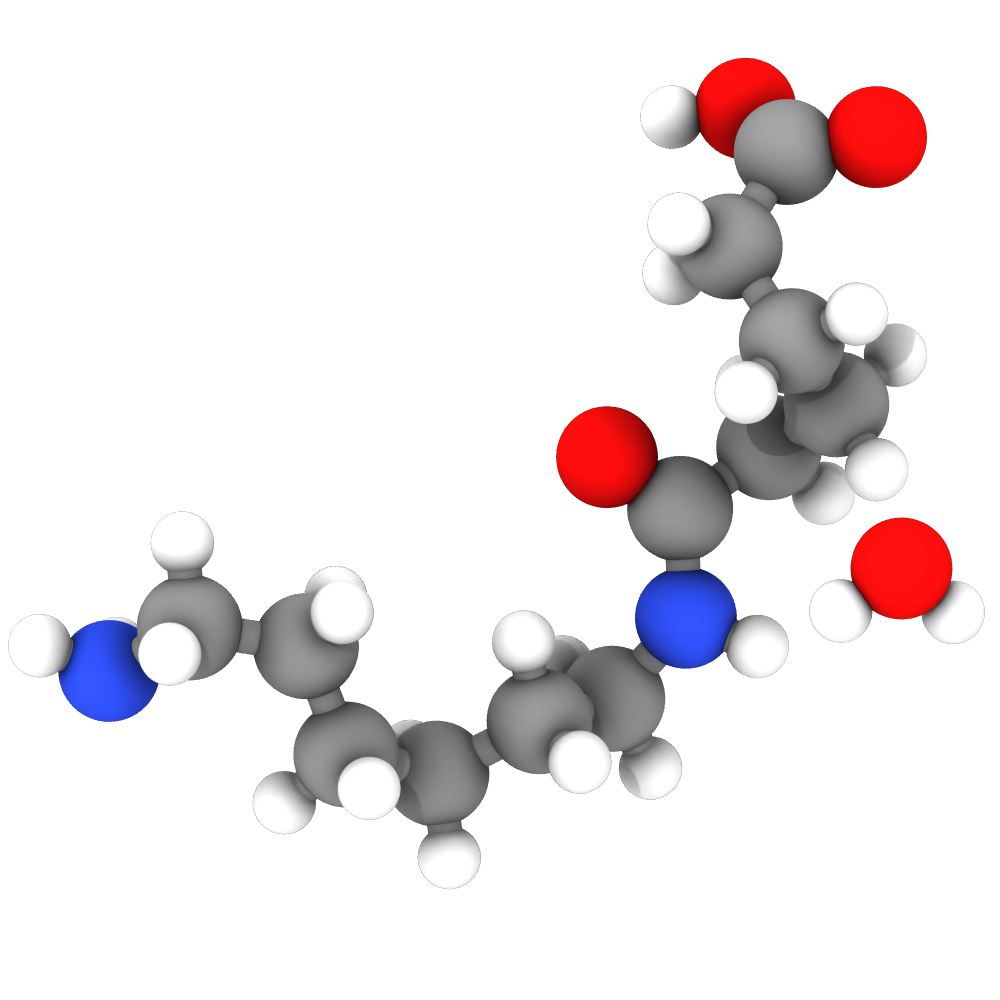
Standard material
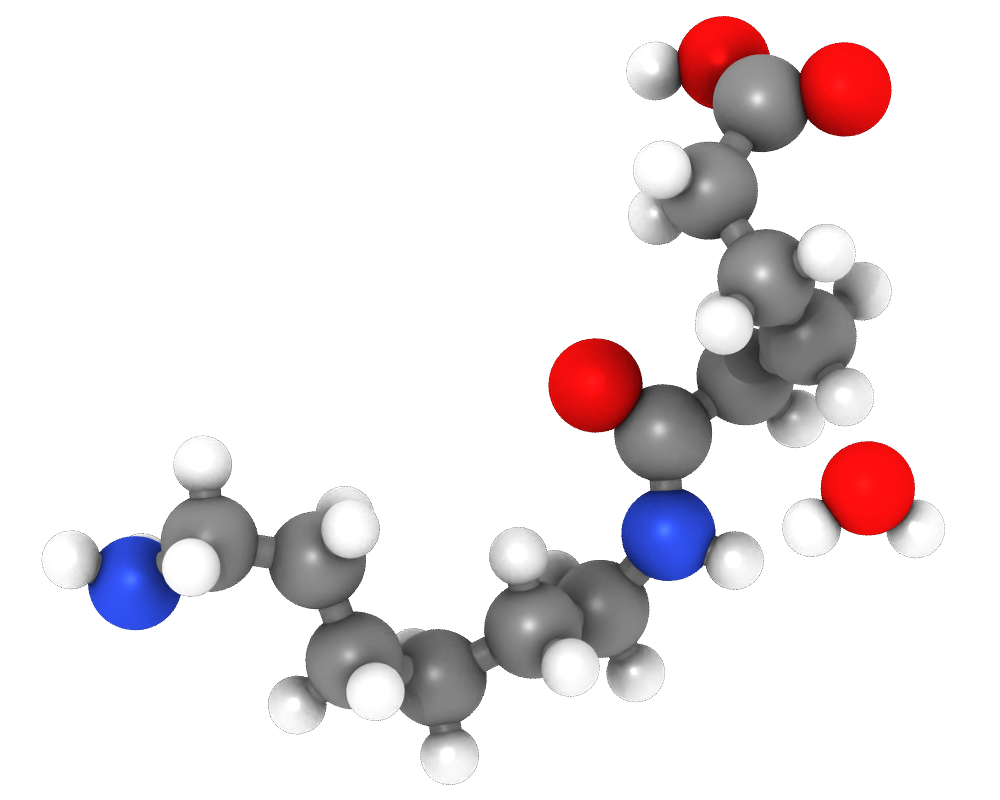
Physically-based material
Furthermore, we added support for depth-of-field rendering (focal blur) effect to the VisRTX renderer.
We’ve added a new color gradient to OVITO that resembles ParaView’s ‘fast’ color map:
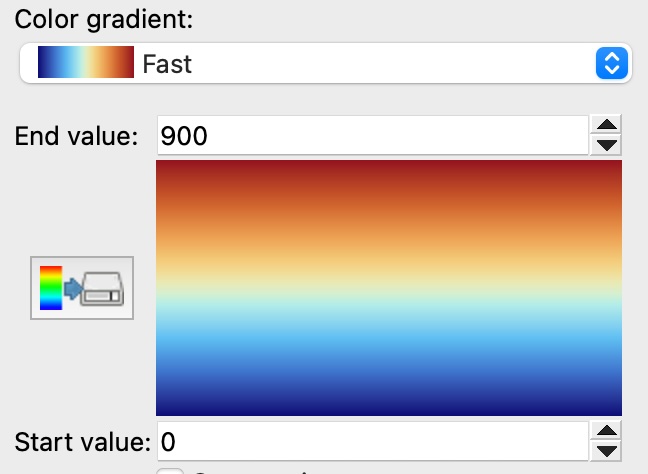
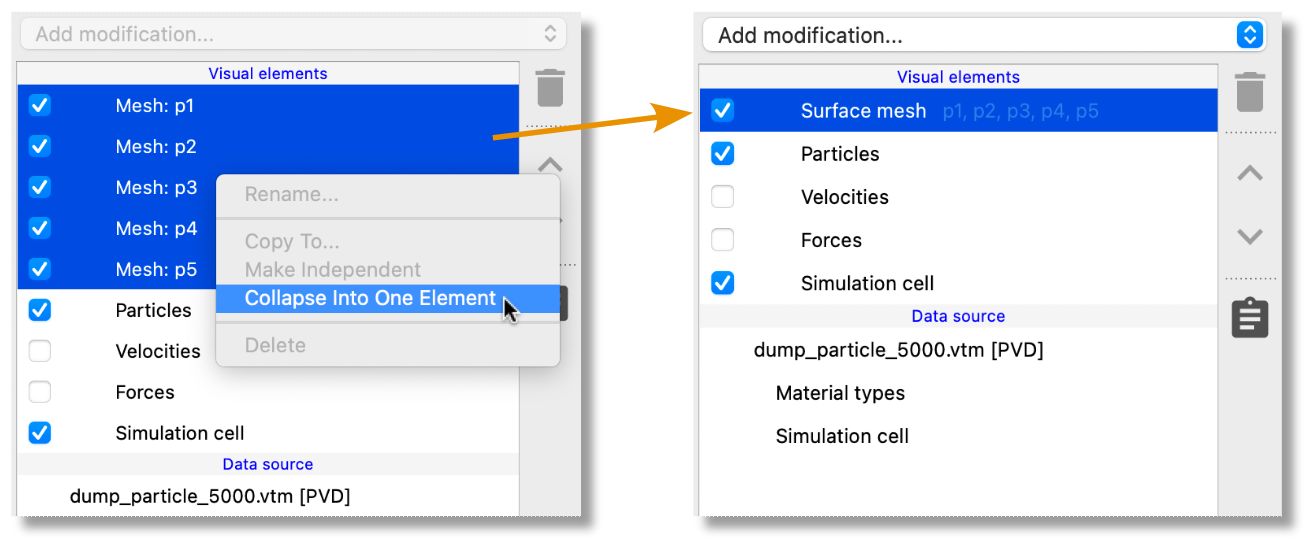
The pipeline editor of OVITO now provides a new function for creating shared visual elements, allowing easy synchronization of visual settings across multiple objects:
When you import a file not containing simulation cell information, OVITO no longer generates an axis-aligned bounding box around the particles. That means, OVITO will operate without any simulation cell in such cases. However, we’ve added the Generate bounding box if needed option to the file reader settings, which allows you to restore the previous behavior if needed.

LAMMPS data file reader: added support for atom styles spin, sph, rheo, rheo/thermal, and bpm/sphere.
A redesigned pipeline status widget, which can now display the complete status text in a tooltip window during mouse hover.
Fix: Missing last tick mark in the color legend overlay.
Fix: Usage-based sorting of “Quick command search” list not working.
Added file export function to Surfaces tab of the data inspector.
Renamed export format “Table of Values” to “Table of Global Attributes” for clarity.
Aspherix PVD file reader: reads simulation time as global attribute
Timeinstead ofTimestep.
Version 3.13.1 (08-Aug-2025)
Radial distribution function (RDF) modifier: Can now break down the computed coordination numbers into different particle types, which is useful for analyzing the local neighborhood’s chemical composition
LAMMPS data file reader/writer: Added support for atom styles spin, sph, rheo, rheo/thermal, bpm/sphere
GALAMOST file reader: Added support for
<force>and<virial>tags and graceful handling of unknown tags in the XML fileVTK XML file reader: Added support for reading surface meshes extracted by ParaView’s Extract Surface filter
Improved rendering performance of the
ovito.data.Linesvisual elementFix: LAMMPS dump file exporter outputs invalid general triclinic simulation cell info
Recognition criteria for binary LAMMPS dump files have been made stricter so that auto-detection of DCD files is not disrupted and misclassification is less likely
pro VisRTX renderer : Users can now choose between the fast standard material and the visually richer physically-based material
pro Added usage example to Spatial Binning modifier documentation, demonstrating how to compute the local stoichiometry of a particle system
Version 3.13.0 (03-Jul-2025)
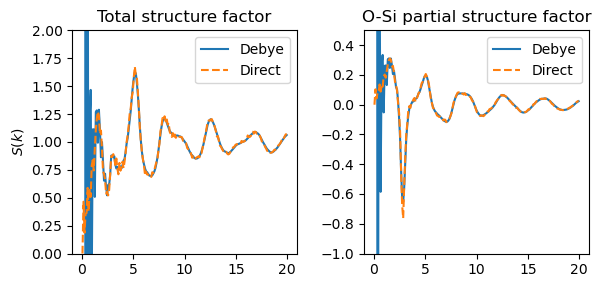
pro New modifier for calculating the structure factor of a particle system:
The Histogram modifier received a new option for selecting the bin normalization mode:
Absolute count: Output the number of elements in each bin.
Relative frequency: Normalize the bin counts to the total number of input elements. Useful for comparing histograms with different numbers of input elements.
Probability density: Normalize the bin counts to the total number of input elements and the bin width. This option is useful for comparing histograms with different bin widths.
pro The OSPRay renderer and VisRTX renderer now can do volume renderings of voxel grids, which lets you visualize volumetric data such as electron density or coarse-grained particle property fields. For this, the Voxel grid visual element has been extended with a new Volume representation mode and an editor for the opacity transfer function:
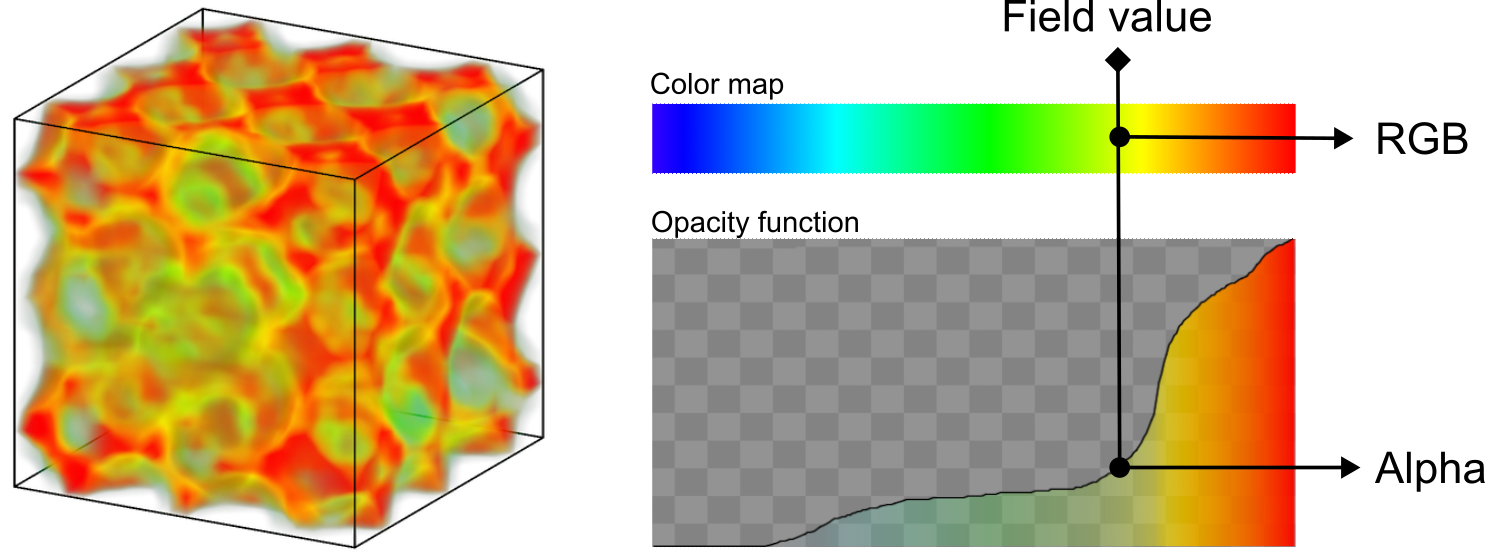

pro The OSPRay renderer now lets you choose a material shading model. The new Principled material implements a physically-based shading model for scene objects, which allows for more realistic rendering of materials with different surface properties (e.g. plastic, metal, glass):
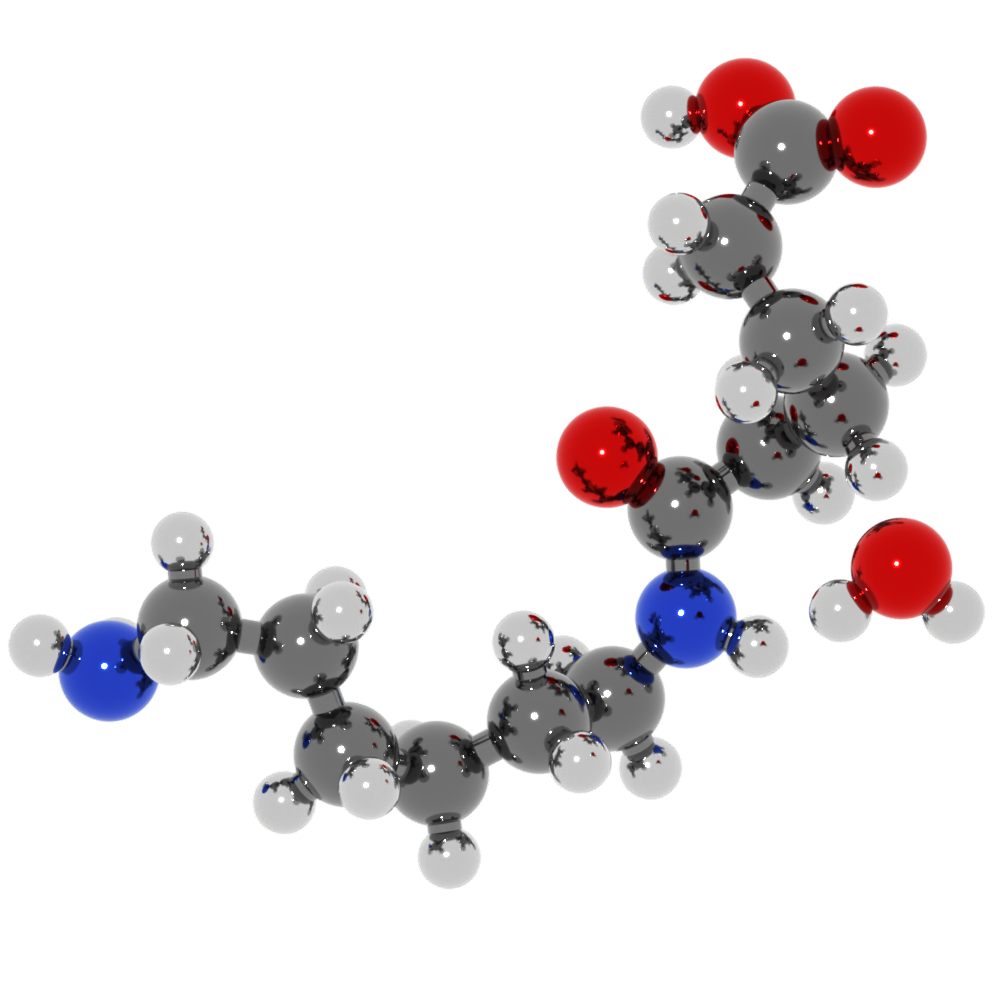
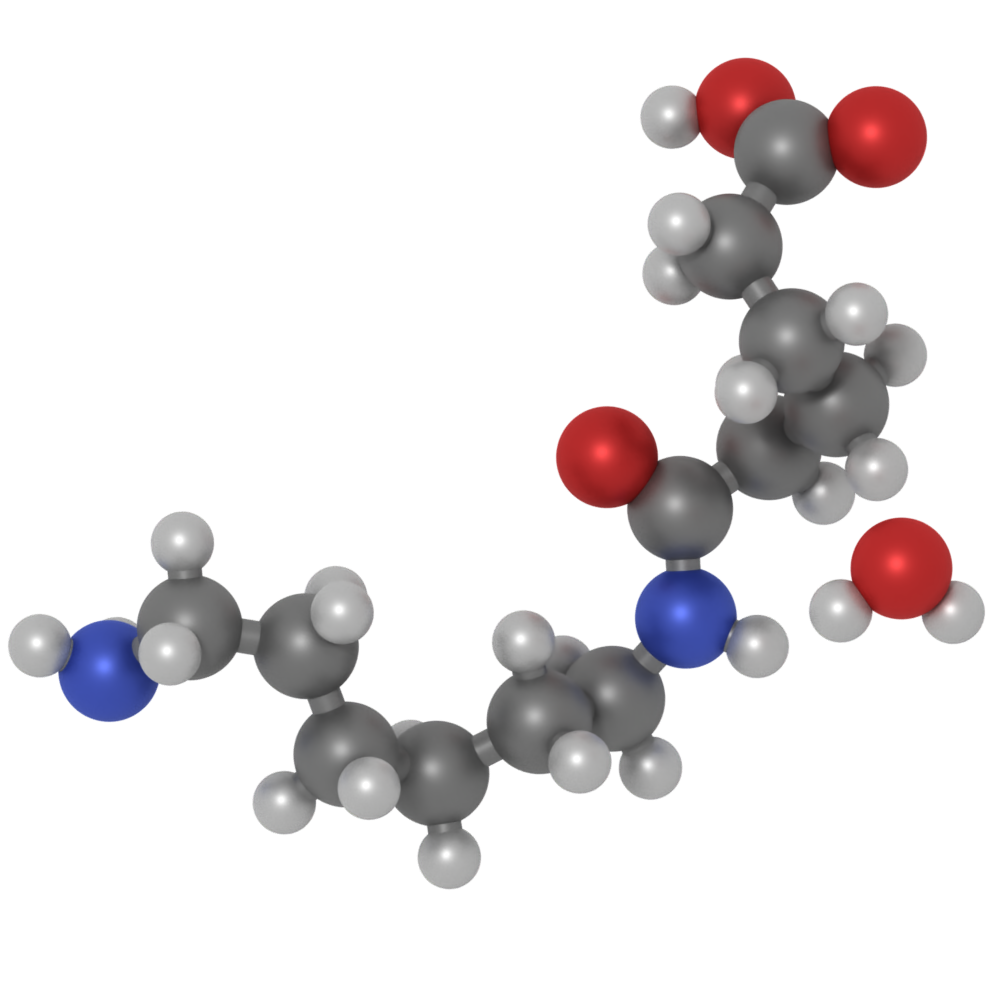
pro The VisRTX renderer now uses a physically-based shading model for scene objects, which also allows for more realistic rendering of materials with different surface properties (e.g. metallic materials).
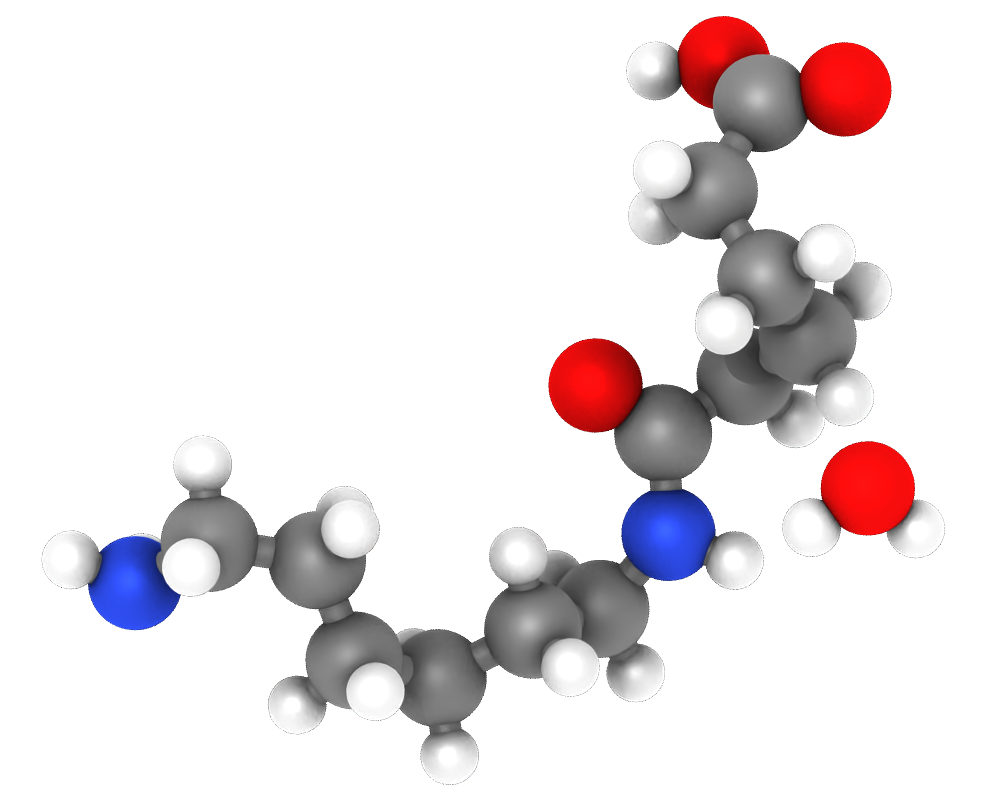

The Freeze Property modifier can now gracefully handle simulations with varying number of particles. It provides a new option to accept unknown particles that were not present in the initial simulation frame.
Aspherix file reader: Added support for data arrays with vector component names
Quantum Espresso file reader: Support more flexible file syntax (see forum discussion)
Fix: Delete selected modifier not working for multiple Lines or Vectors objects in the data collection.
Global attribute identifiers starting with a
.are now hidden in the data inspector panel.pro Cleanup redundant code output for visual elements in the Python code generator
py The
Viewport.render_anim()method now gives you more control over the generated filename sequence when rendering an image file series. The wildcard character'*'in the filename parameter gets replaced with the current numeric frame number:vp.render_anim("image_*.png", size=(640,480))py Added
return_distancesparameter toovito.data.SurfaceMesh.locate_point()method.py Added
__hash__method to OVITO object classes, which allows using OVITO objects as dictionary keyspy Fixed a locale issue on Linux systems where the
LC_*environment variables are set to a non-English locale.py Added preliminary API for computing 3D Delaunay tessellations:
ovito.data.DelaunayTessellation
Version 3.12.4 (26-May-2025)
pro Fixed OSPRay library loading error on Linux:
[ERROR] Load of ospray_module_denoiser failedLines objects: Fixed hover information display in status bar for PBC wrapped line segments
Fix: (very rare) bug in 2d polygon tessellation code used by the surface mesh visual element, which could lead to an infinite loop in pathological cases
Version 3.12.3 (12-May-2025)
Added support for the GROMACS TRR file format
Added support for the MercuryDPM trajectory data format
Fix: Affine transformation modifier refuses to apply a translation in addition to a rotation to particles with the
OrientationpropertyFix: Compute property modifier: Named types in expressions not working for property variables starting with ‘@’
Fix: Vectors visual element crashes if the input property has an unexpected data type or component count
pro Upgraded embedded Python interpreter to Python 3.12.10 (Linux) and Python 3.13.3 (Windows)
Version 3.12.2 (23-Apr-2025)
The file column mapping dialog now includes a context menu function for toggling the checked state of multiple columns at once
Bug fix: Crash in the Particles tab of the application settings dialog when using the “Restore built-in defaults” function (OVITO 3.11.0 regression)
py Bug fix: Writes to a NumPy view of the
Selectionproperty array, followed by an application of theDeleteSelectedModifiermodifier, may lead to wrong results (OVITO 3.11.0 regression)pro Suppress unnecessary Python code generation for
ComputePropertyModifier.neighbor_expressionsfield if modifier is not set to operate on particles (OVITO 3.12.0 regression)
Version 3.12.1 (02-Apr-2025)
Fixed visual glitch when moving the cursor into the 3D viewport window, which occurred on some Windows/Linux systems in OVITO 3.12.0
Compute property modifier: Now allows creating user-defined vector properties
Compute property modifier: Added the special variables
SpatialPositionandVoxelCoordinatefor voxel grid computationsColor coding modifier: Added new color gradient “Cyclic Rainbow”, which is useful for visualizing cyclic properties such as phase angles or in-plane orientations
Color legend layer: Fixed reordering of DXA dislocation types list during trajectory playback
Dislocation analysis (DXA) modifier: Fixed crash in function “Mark dislocation core atoms” when no dislocations are present
Voronoi analysis modifier: Redesigned UI
Fix: GUI function “Reset to default” not working for numerical parameter fields
py Extended the
ComputePropertyModifier.expressionsparameter to support the creation of user-defined vector properties
Version 3.12.0 (26-Feb-2025)
A new command panel tab, which can host various utility applets (e.g. Python scripts, custom tools, or the existing remote rendering function)
Support for type names in selection expressions:

Compute property modifier: New option to include all bonded neighbor particles in the computation:

The LAMMPS dump local file reader in combination with the Load trajectory modifier now supports loading dynamic angle/dihedral/improper interactions from LAMMPS dump local files. In addition, it can now automatically recognize the columns of the dump file if you follow the instructions given in Automatic column mapping.
OVITO can now read a zstd compressed trajectory file while new frames are being appended to it by the running simulation code. For gzipped and uncompressed trajectory files, this already worked.
GSD/HOOMD file reader: Retain complete chunk paths in per-particle property names loaded from GSD files
GSD/HOOMD file reader: Automatically generate Vectors visual elements for user-defined particle properties with 3 components
Aspherix file reader: Fixed grid domain size incorrectly overriding the simulation cell size
Restored automatic seeking to the last frame of a growing trajectory file when using the update trajectory frames function
py The new
ovito.io.FileWriterInterfaceclass allows writing custom file exporters for OVITO Pro and the OVITO Python module:class MyFileWriter(FileWriterInterface): def write(self, *, filename, frames, pipeline, **kwargs): with open(filename, 'w') as file: file.write("...")
export_file(pipeline, "output.dat", format=MyFileWriter)
py Added the new
ovito.guimodule, which contains GUI-related functions, e.g., functions for creating visible viewport windows.py Added the
ovito.gui.UtilityInterfaceextension class and theovito.traits.action_handler()decorator, which allow defining custom action handlers for UI push buttons, e.g., in custom utility applets:class MyUtilityApplet(UtilityInterface): export_btn = Button(ovito_label="Export my data") @action_handler("export_btn") def export_data(self): pipeline = scene.pipelines.selected_pipeline export_file(pipeline, "data.txt", "txt/attr", columns=["CommonNeighborAnalysis.counts.FCC"])
py Added translation and rotation parameters to
Pipeline.add_to_scene()and removedPipeline.translationandPipeline.rotationpropertiespy Replaced
LinesVis.shadingparameter withLinesVis.flat_shadingparameterpy Replaced
DislocationVis.shadingparameter withDislocationVis.flat_shadingparameterpro Added OVITO Pro Python extensions gallery, which provides an easy way to discover and install third-party Python extensions:

pro Added the new ASE trajectory writer
pro Added extension interface for custom file exporters
pro Added extension interface for custom utility applets running in the command panel of OVITO Pro
pro Added a GUI function for easily installing third-party Python packages in the embedded interpreter of OVITO Pro:


pro OVITO Pro can now be used as a Jupyter kernel (in conda environments), combining interactive Python scripting with the full OVITO Pro GUI:
pro New depth-aware outline effect in OSPRay renderer and VisRTX renderer :


pro Improved object picking performance in interactive viewports for the VisRTX renderer
Version 3.11.3 (29-Dec-2024)
Automatically insert the Create isosurface modifier into the pipeline when importing a CHGCAR file
GSD file reader: Added support for sphere and spherocylinder particle shapes (ConvexPolyhedron with 1 or 2 vertices)
Workaround for Qt/OpenGL memory leak on macOS when using the Ambient occlusion modifier
py Added
pymatgen_to_ovito()andovito_to_pymatgen()functions for converting OVITO particle structures to and from pymatgen Structure objectspy Notify user of invalid usage of
ovito.data.CutoffNeighborFinderiterators (matsci.org discussion)
Version 3.11.2 (28-Nov-2024)
Affine transformation modifier: (pure) rotations are now applied to the
Orientationproperty of particles if presentRadial distribution function (RDF) modifier: Exclude types of unselected particles from partial RDF legend if Use only selected particles option is enabled
Fixed crash when closing OVITO while Generate trajectory lines modifier is still processing
Fixed GUI issue on Linux: Pressing enter key while focus is in a text field may toggle a parent group checkbox
py Added the
ovito.traits.Matrix3parameter trait typepro VisRTX: Performance improvements for rendering particles with user-defined shapes
pro VisRTX: Fixed program crash when rendering multiple interactive viewports in parallel
pro VisRTX: Fixed color compositing of semi-transparent objects in interactive viewports
pro VisRTX: Reduced z-fighting artifacts along wireframe lines in interactive viewports
pro VisRTX: Improved object picking speed in interactive viewports for large particle datasets
Version 3.11.1 (11-Nov-2024)
Added read/write support for zstandard (*.zst) compressed files (e.g. custom/zstd LAMMPS dump style)
New Viewport Graphics Configuration dialog
Added support for VTK files with header type UInt32
Fixed regression: Identify diamond structure modifier reports all atoms as “OTHER”
py Added support for Python 3.13
py Windows: Ensure that DLLs are always loaded from the package directory and never from other locations in the Windows search path
Version 3.11.0 (08-Oct-2024)

Visualization of vector information
We’ve added the Vectors object type to OVITO and the corresponding ovito.data.Vectors Python class,
which allow placing arrow glyphs at arbitrary locations in 3d space (independent from particles) to visualize vectorial information anywhere.

Symmetric range option for color mapping tools

The Color coding modifier and other color mapping tools can now maintain a symmetric value range, which is useful for visualizing scalar fields that can take both positive and negative values. If this option is enabled, the color mapping range is automatically centered around zero.
Template system for viewport layers
Similar to modifier templates, you can now create pre-configured templates for viewport layers and reuse them in future program sessions. This feature is particularly useful for creating consistent overlays across multiple visualizations.
Interactive viewport rendering with VisRTX pro
On Windows and Linux machines with a CUDA-capable NVIDIA GPU, OVITO Pro now supports interactive rendering with the high-performance VisRTX ray-tracing backend. This allows you to explore your data interactively in the viewports with high-fidelity rendering quality.
Demo versions of the high-fidelity rendering backends in OVITO Basic
OVITO Basic now includes demo versions of the rendering backends OSPRay, Tachyon, and VisRTX.

Calculation of dislocation density and Nye tensor fields from DXA output pro
We’ve extended the Spatial binning modifier to compute the dislocation density field from the output of the DXA analysis. The new option also supports calculating the Nye tensor field from the discrete dislocation lines.
Identification of dislocation core atoms pro

We’ve developed a method for identifying atoms that are part of the defect core of individual dislocations. This pro feature is now available in the Dislocation analysis (DXA) modifier as a new option, see Identification of dislocation core atoms.
Python code generator for complex pipeline architectures pro
The Python code generator now supports scenes containing multiple pipelines and can generate valid script code for the most complex pipeline architectures, including branched pipelines, shared modifiers, and shared visual elements created with the interactive OVITO GUI. It also got better at handling viewport layers in complex visualization setups.

Improved rendering quality for semi-transparent objects and antialiased edges pro
OSPRay renderer : Improved rendering quality for semi-transparent objects and fixed dark artifacts along object edges on light backgrounds

Old

New
Further improvements in this program release:
Major code refactoring to improve performance and maintainability; elimination of unnecessary data copies in the pipeline system
Modifiers now keep their computed results in memory when being temporarily turned off by the user to avoid recomputation when reenabled again
Better decoupling of UI and pipeline system, in particular to speed up the processing of long trajectories, which caused an unnecessary number of display updates in the past
Generate trajectory lines modifier: Trajectory processing now happens automatically in the background - it is no longer necessary to start the process manually
Affine transformation modifier: Rotational transformations now act on the Burgers vectors of dislocation lines
Dislocation analysis (DXA) modifier: Ensure cross-platform reproducible output
Unwrap trajectories modifier: Performance optimizations
Added a notification dialog that is shown on application startup when new program updates become available
Simulation file import: Updated mass of Zn in internal table of elements from 65.409 (pre 2007 value) to 65.38 (see https://www.ciaaw.org/zinc.htm) - old value is still recognized for compatibility reasons
The Aspherix file reader can now load both particle-particle and particle-wall contact networks
Added parser support for modern Aspherix .vtm files containing a “Bodies” section. Bodies information from non-convex particle simulations is read in as a data table.

Each Aspherix DEM body is originally composed of multiple convex sub-particles.

A Python extension allows combining the sub-particles to non-convex body particles.
Import of existing CA files (i.e. loading of precomputed DXA results) is now an exclusive OVITO Pro program feature
Slice and Affine transformation modifiers: Step-wise parameter increments are now proportional to the simulation box size instead of the current parameter value to improve usability during interactive adjustments
Trailing “_” in imported particle property names are now stripped to avoid possible conflicts with the underscore notation in the Python API
Updated third-party components to latest versions: Python, OpenSSL, Qt, PySide6, Qwt
pro Time averaging modifier: Added time-averaging of the simulation cell shape and new option to overwrite original values with average
pro OSPRay renderer : Added support for pseudo-color mapping
pro VisRTX renderer : Improved rendering performance for scenes with large numbers of cubic, ellipsoidal, or superquadric particles
pro Environment variable
OVITO_SAFE_MODE=1can be set to effectively block execution of Python scripts embedded in .ovito session state files from untrusted sourcespro Updated the standard code template in the Python script modifier to use the Advanced programming interface
Python API additions and changes:
py Added support for NumPy 2.x
py Full compatibility with Python 3.12
py The OVITO module now initializes the global Qt application object only on demand to avoid conflicts with other Python packages that also use PySide6
py
ConstructSurfaceModifier: Optionmap_particles_to_regionsnow outputs per-region particle membership lists- py The
ovito.data.SurfaceMesh.locate_point()method has been vectorized and can now process an array of input points using all CPU cores py New method
NearestNeighborFinder.find_all_at()to efficiently determine the closest particles around several spatial locations at oncepy New method
SimulationCell.wrap_point()to map one or more points into the primary simulation cellpy New attribute
ovito.pipeline.Pipeline.num_frames, which allows querying the number of (output) trajectory frames of a pipelinepy New attribute
ovito.pipeline.Pipeline.frames, which allows iterating over the output data collections computed by a pipeline for all trajectory framespy New attributes for file source objects:
py New advanced methods:
py New attributes in class
ViewportOverlayInterface.Canvas:py New parameter trait types:
py New class
ovito.data.DataObject.Refand methodsovito.data.DataCollection.get()andovito.data.PropertyContainer.get()py
ColorLegendOverlay: Addedpipelinefieldpy Added the
HistogramModifier.select_elementsoptionpy Added support for Python’s copy module, which allows creating exact copies of OVITO data objects
py Deprecated method
GenerateTrajectoryLinesModifier.generate()(trajectory line generation is now done automatically by the modifier)
Bug fixes:
Data table plots: Axis scales and labels are invisible (white on white) in exported plots if dark UI theme is active
Data table plots: Regression due to updated Qt framework: missing colors in plot legends
Mouse cursor wrapping in spinner widgets not working for vertical multi-screen setups
Mouse cursor wrapping requires accessibility access on macOS
pro Time averaging modifier uses wrong divisor in average calculation if trajectory length is not an integer multiple of the sampling frequency
pro Render LAMMPS regions modifier: unexpected error if the modifier is inserted more than once into the same pipeline
Version 3.10.6 (03-May-2024)
LAMMPS data and dump files: Added I/O support for general triclinic simulation cells, which will soon be introduced by LAMMPS
LAMMPS dump and IMD file export: Use correct vector component names in output file columns for user-defined particle properties
py
ovito.io.ase.ase_to_ovito()function: filter out non-string metadata keys if present in the ASE Atoms objectpy
ovito.io.import_file()function: Use Python’s getpass() function to prompt for SSH password without echoing it to the console
Version 3.10.5 (17-Apr-2024)
Added new standard particle property
Vector Transparency, which allows controlling the transparency of vector glyphs on a per-particle basisFixed endless loop when trying to cancel SSH authentification dialog after opening an existing session state file
Fixed a bug that prevented user changes to particle type parameters in combination with the Unwrap trajectories modifier
Radial distribution function (RDF) modifier: Lifted upper limit on the number of RDF bins
py Added option
OpenGLRenderer.order_independent_transparencypy Documented the capability of
ovito.io.import_file()to load files from remote SSH and HTTPS serverspro Corrected sun-sky light brightness scale of OSPRay renderer to match old behavior (v3.10.0 regression)
Version 3.10.4 (13-Mar-2024)
Adjust view dialog: Added option to numerically control the camera’s roll angle
Updated third-party libraries: OpenSSL 3.0.13, libssh 0.10.6, ffmpeg 6.1.1, Python 3.11.8, VisRTX 0.8.0
py Added
CreateIsosurfaceModifier.smoothing_levelandCreateIsosurfaceModifier.identify_regionsoptionspro Create isosurface modifier: New option to identify spatial regions enclosed by the isosurface
pro Improved glTF file export: particles and bonds may now be exported as single meshes for better rendering performance (but larger file size)
pro OSPRay renderer : Fixed picking of focal length in the interactive viewports
pro VisRTX renderer : Added mesh backface culling support, gracefully handle initialization errors
pro Construct surface mesh modifier: Changed how the “external” spatial region is defined in open (non-periodic) systems, now having its volume reported as
inf
Version 3.10.3 (19-Feb-2024)
Fix: Lines visual element renders wrong line caps when using option “Show up to current time only”
Fix: Drop-down list of available modifiers inserts wrong modifier template into pipeline after an entry was added to the list
py Fix:
create_jupyter_widget()method failing (issue #229)py Fix: Issue with
ovito.traits.Colornot accepting a NumPy arraypro Added parameter for ambient occlusion cutoff to VisRTX renderer
pro Fix: Warnings in Render LAMMPS regions modifier due to code line duplications
Version 3.10.2 (02-Feb-2024)
Rendering of animated GIFs with transparent background color
LAMMPS data file exporter: Emit 0 atom/bond types if no types are defined and there are no particles/bonds
Load trajectory: Drop ‘Periodic Image’ particle property from topology dataset if trajectory file does not contain dynamic image flags
py Added Python API for accessing the line connectivity information in a
DislocationNetworkpro Fixed OSPRay image rendering on transparent background (v3.10.0 regression)
pro macOS: Fixed crash in OSPRay renderer due to missing dylib (v3.10.0 regression)
Version 3.10.1 (09-Jan-2024)
Added support for the LAMMPS dump yaml file format to the LAMMPS dump file reader
Replicate modifier: Use particle property
Periodic Imageif present to yield correct replicated molecule identifiersFix: Frame 0 of LAMMPS dump, xyz and pdb trajectory files gets loaded a second time unnecessarily
py New Python properties
Pipeline.translationandPipeline.rotation, which control the placement of a pipeline’s visual output in the 3d scenepy Fixed offscreen font rendering in standalone Python module on (headless) Linux platform
Version 3.10.0 (28-Dec-2023)
Perspective distortion for axis indicators
The Coordinate tripod layer provides a new option to distort the displayed tripod according to the perspective projection. Now the axes indicators exactly align with the edges of the simulation cell:
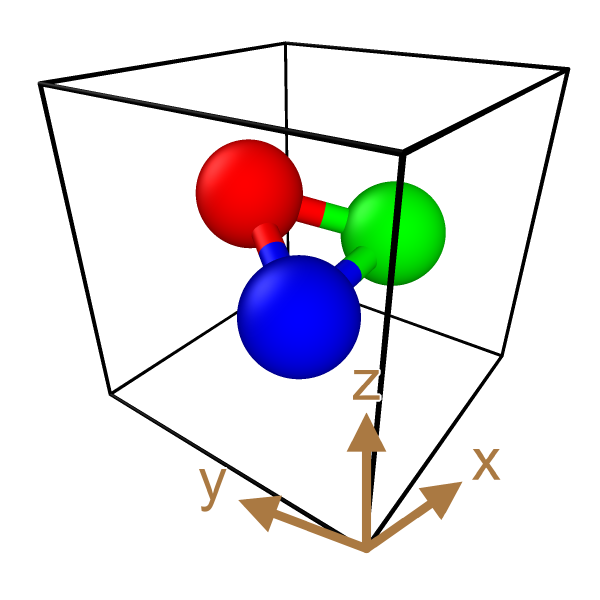
OVITO 3.9
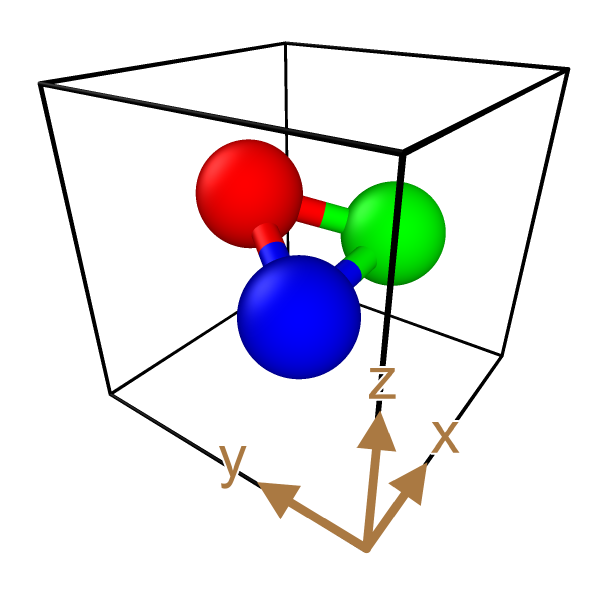
OVITO 3.10
3D full-scene export to glTF format pro
The new glTF file exporter can export OVITO models including all visual elements. This lets you export entire scenes to PowerPoint, Blender, or the web as movable 3d objects:
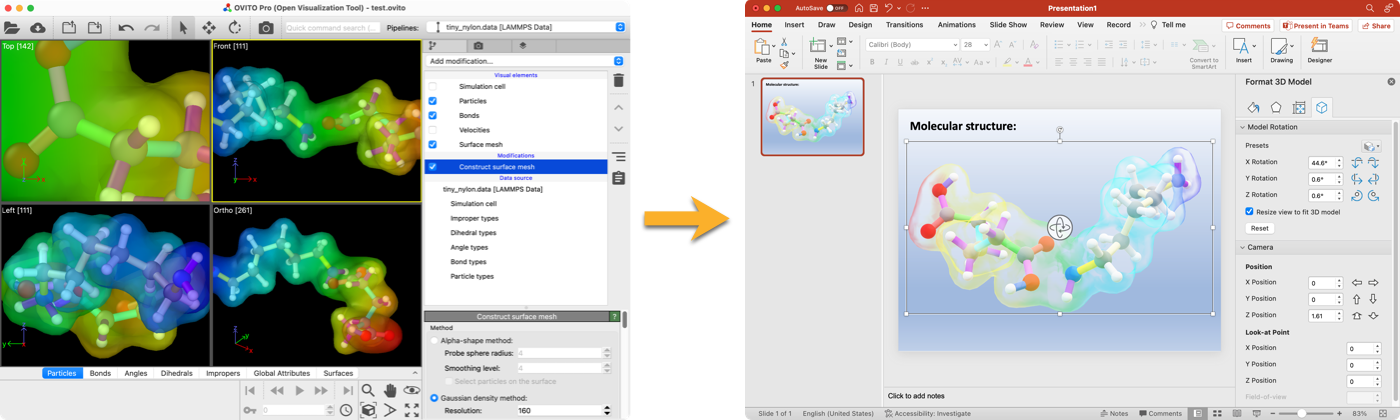
Below is a glTF model exported from OVITO Pro, embedded into this HTML page. Click and drag to rotate the model.
The 3d viewer works only in the online version of this document due to web browser security restrictions.
In the offline version of the OVITO docs, you will see only a static image of the model.
New Python classes to paint arbitrary 3d lines pro
The new ovito.data.Lines and ovito.vis.LinesVis classes allow
visualizing line segments in 3d space, e.g., for augmenting particle models with additional
information.
pro New renderer: NVIDIA VisRTX
We’ve added the VisRTX renderer and a corresponding Python class ovito.vis.AnariRenderer,
which make use of the cross-vendor Khronos ANARI API.
The VisRTX rendering backend offers hardware-accelerated ray-tracing on NVIDIA GPUs and can generate high-fidelity scene renderings
in a fraction of a second – even for complex datasets containing millions of objects.
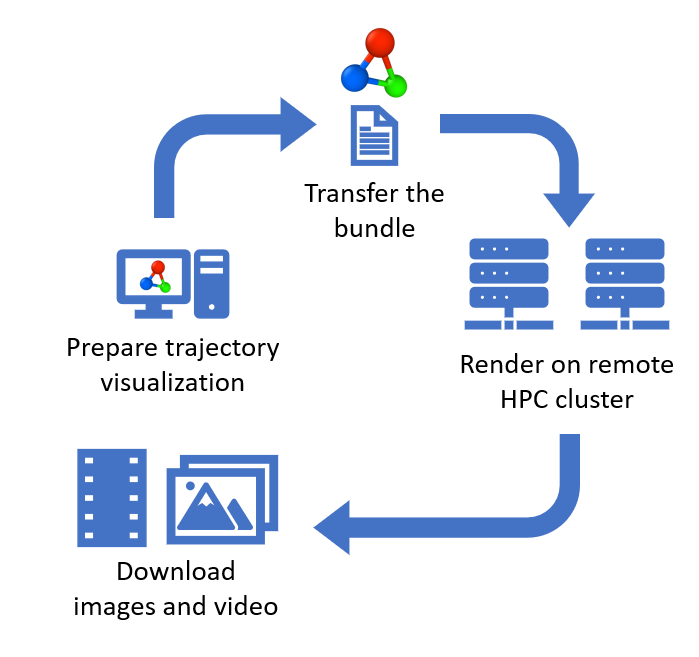
Remote trajectory rendering function pro
Added a function for Rendering simulation trajectories on remote computers . It lets you prepare trajectory visualizations on your local computer and easily render them on a parallel HPC cluster. OVITO Pro takes care of packaging all required data files and generating the necessary job scripts for you.
Further changes in this program release:
Smooth trajectory modifier: Now supports trajectories with varying number of particles
Smooth trajectory modifier: Fixed wrong interpolation/averaging of particle orientations
LAMMPS data file reader: Tolerate more than one empty line after file section titles
XYZ file reader: Automatic detection of reduced coordinates turned off by default, because extended XYZ files with reduced coordinates are very rare
py Some Python functions now return true NumPy arrays instead of Python tuples
py New Python function
DislocatioNetwork.Line.point_along_line()py New parameter trait types
ovito.traits.FilePath,Vector2, andVector3py Renamed existing parameter traits types
ovito.traits.OvitoObjectandovito.traits.Colorpy Restricted
ovito.Scene.load()to session state files written by OVITO Pro or the Python modulepy Added function parameter
pipeline_nodetoModifierInterface.modify()py
SurfaceMeshTopologyclass now performs out-of-range checks on function parameters.pro OpenGL, OSPRay, and Tachyon renderers: Added buttons to reset numeric parameters to their default values
pro User-defined parameters can now be grouped in the UI by means of the new
ovito_groupmetadata attribute
Version 3.9.4 (04-Nov-2023)
Fix: OpenGL rendering of ellipsoidal/superquadric/box particles with wrong orientations (regression since v3.9.0)
Version 3.9.3 (01-Nov-2023)
Expand selection modifier: Added new output attribute ExpandSelection.num_added
Identify diamond structure modifier: Added missing output attribute IdentifyDiamond.counts.OTHER
Fix: Section “Modifier templates” of available modifiers list not updated correctly when adding/removing templates
Fix: LAMMPS data file reader: vector vis settings of Velocity property lost after loading a state file
Fix: Segfault when opening a .ovito state file with macOS Finder while OVITO is already running
Fix: Construct surface mesh modifier: Option Map particles to regions may yield invalid results when used with option Use only selected input particles
Updated third-party components: OpenSSL 1.1.1w, Qt/PySide6 6.5.3, Python 3.11.6
py PyPI packages for Python 3.12
Version 3.9.2 (31-Aug-2023)
Support Ctrl+C copy to clipboard in table of distances, angles, and dislocations in the data inspector
Fix: Text label viewport layer accidentally disables 3d depth test in interactive viewports if used as an underlay
py
render_image()andrender_anim()now raise exceptions in case an error occurs in any of the scene pipelines (can be changed via new parameter stop_on_error)py New flag
Pipeline.preliminary_updatespy Corrected data column headers in XYZ, LAMMPS dump, and IMD files written via
export_file()if a vector property was specified in the columns listpro New class-based programming interface for custom viewport overlays:
ovito.vis.ViewportOverlayInterfacepro Build Conda package as monolithic binaries for improved performance of the Python interface
pro Updated third-party components: OpenSSL 1.1.1v, PySide6 6.5.2, Python 3.11.5
Version 3.9.1 (06-Aug-2023)
Fix: Voronoi Analysis modifier crashes if simulation cell is degenerate or atom count is zero, and option Generate neighbor bonds is turned on
py New Python class
ovito.pipeline.PipelineSourceInterfacepy New Python method
ModifierInterface.compute_trajectory_length(), which gives user-defined modifiers control over the timeline lengthpy New Python field
Modifier.titlepro Fixed ovitos -m pip install failure for packages that require a build step
Version 3.9.0 (02-Aug-2023)
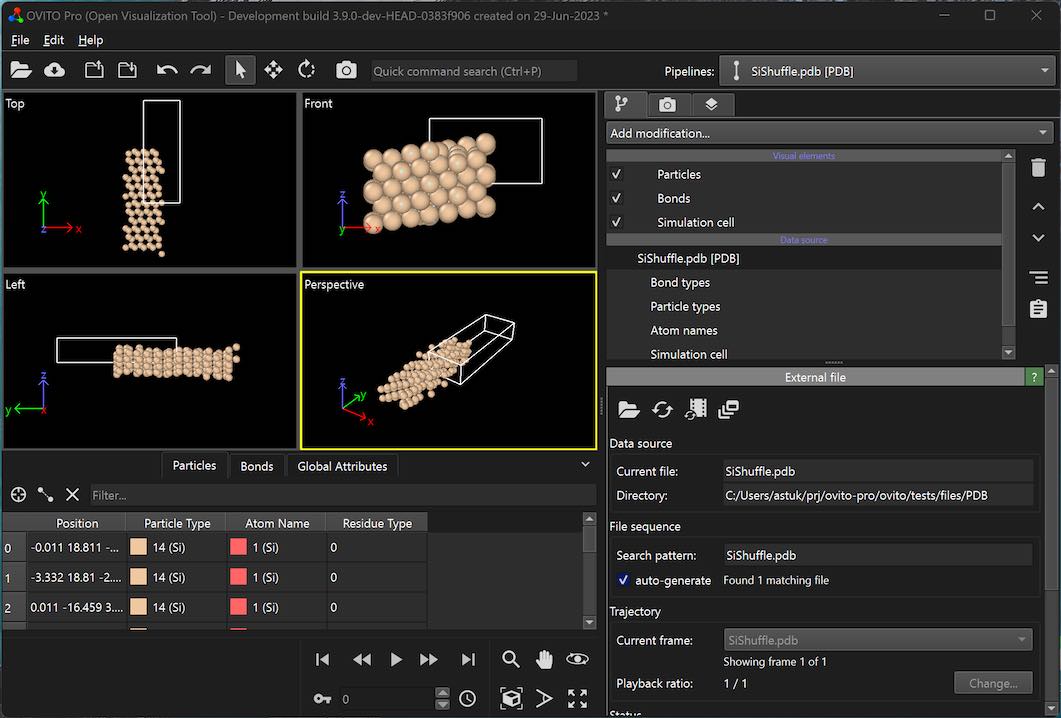
Dark mode support on Windows
To enable the dark UI theme on Windows, go to the application settings and switch on Enable automatic dark mode. OVITO will follow the Windows system color theme.
OpenSSH client integration pro
OVITO Pro is now able to access data files on remote machines using OpenSSH’s sftp utility, which fully supports smartcard authentication and other advanced ssh features. See OpenSSH client for further information.
User-defined file format readers pro
This program release introduces a programming interface for user-defined file readers, which enables
you to develop parser functions for new file formats in Python. User-defined file readers are fully integrated into the
GUI of OVITO Pro and work seamlessly with the import_file() function from the OVITO Python module.
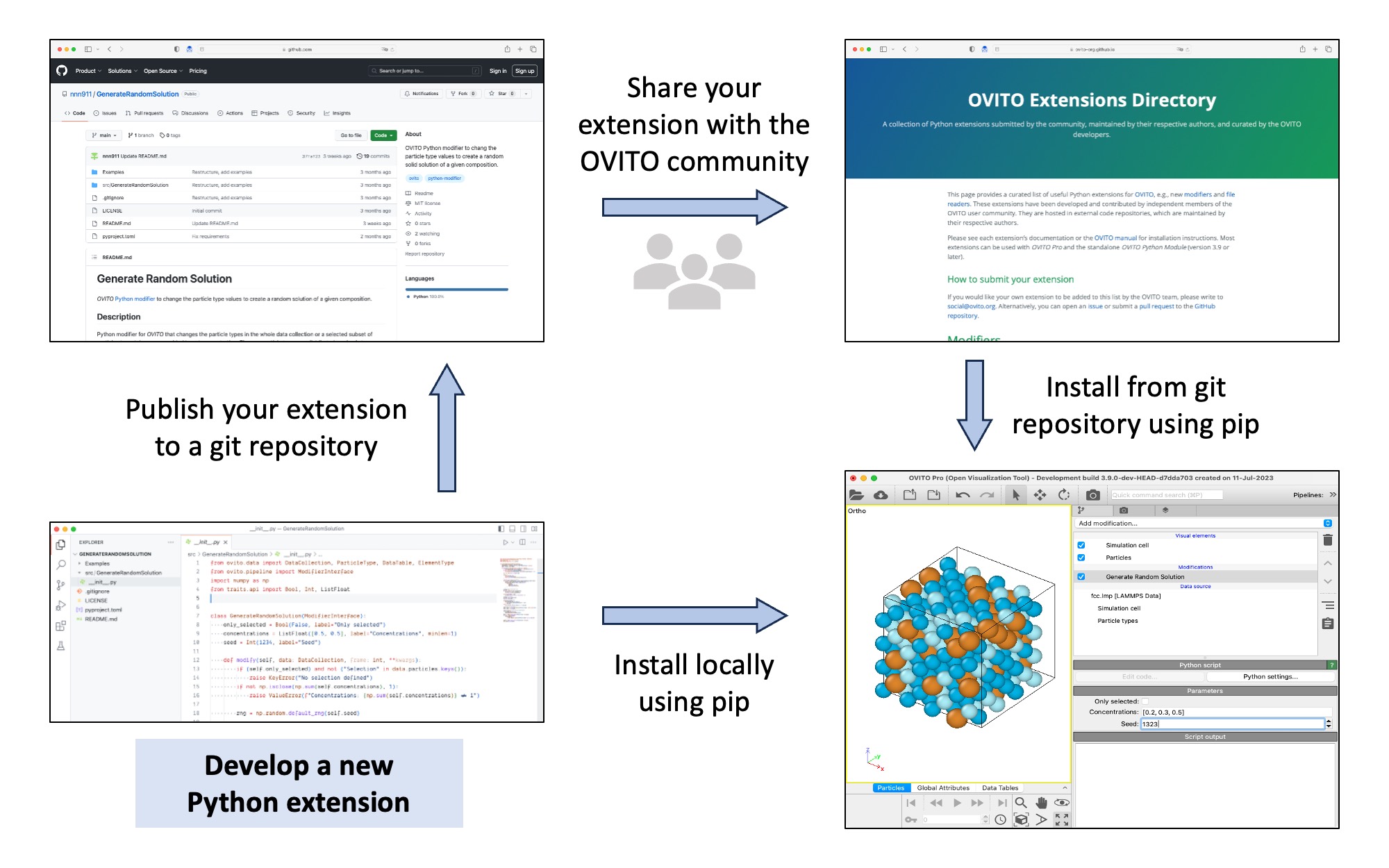
Discovery mechanism for Python extensions pro
Python extensions for OVITO Pro or the OVITO Python module (i.e. user-defined modifiers and file readers) can now be packaged as Python modules, making it easier to deploy and install them (using pip install). Custom extensions you’ve developed can be put under version control in a Git repo and shared online with other OVITO users if desired – we have set up the new OVITO Extensions Directory for that purpose. After easy installation on a user’s computer, OVITO Pro automatically discovers all extensions and makes them available in the GUI.

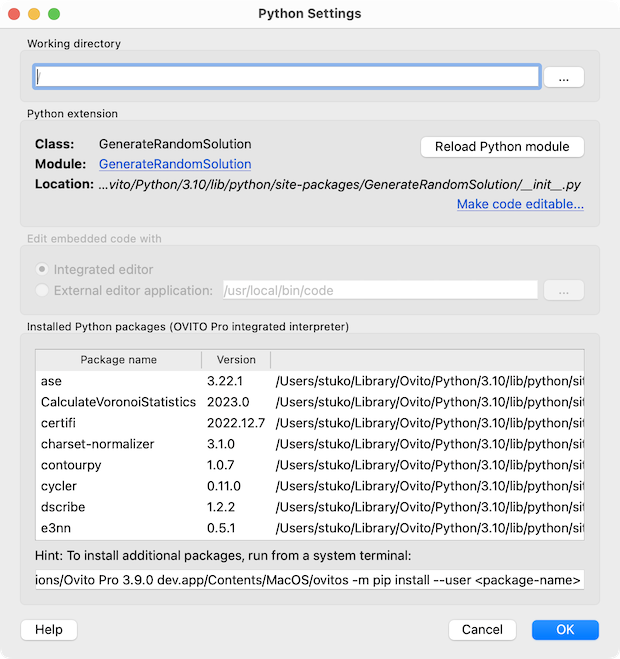
New Python Settings dialog pro
The new Python Settings dialog provides access to all things related to a Python extension in OVITO Pro:
Configure the current working directory used for file I/O operations
Hot reload function for imported Python modules, which streamlines development of Python code located in external source files
Import the source code of installed extensions into the current program session to selectively customize functions when needed
List of all installed Python packages that are available for import by user code
Support for more file formats
OVITO can now import DCD trajectory files, which are written by the CHARMM, NAMD, and LAMMPS simulation codes. OVITO Pro and the OVITO Python module can additionally read ASE trajectory files.
Further changes in this program release:
Support for additional property data types (float32, int8) to reduce memory footprint of particle properties with low precision requirements (e.g. Color, Selection)
OpenGL renderer: Performance optimizations, direct upload of float32 and int8 array values to GPU memory
GSD file reader: Do not skip
log/chunks containing/in their names (issue #226)Fix: Color Coding modifier’s “Adjust Range” function does not follow option “Only selected”
Search patterns for trajectory file series: Avoid asterisk in file extensions containing digits, e.g.
snapshot0000.h5→snapshot*.h5Data table file exporter does not require a
y-property anymoreAutomatic name mangling of atom attributes imported from LAMMPS dump, GSD, and XYZ files in case they do not conform to OVITO’s property naming rules
The Vulkan viewport renderer has been removed
py New Python methods
Property.add_type_idandProperty.add_type_namepy New Python method
VoxelGrid.viewpy Performance optimizations for property data access from Python code
Version 3.8.5 (19-Jun-2023)
Voronoi analysis modifier now outputs per-face
AreaandVoronoi Ordermesh propertiesPDB file reader: Refined detection of cells with periodic boundary conditions
GSD file reader: Support time-varying radii of spherical type shapes; display simulation step numbers in timeline
LAMMPS dump file reader: Automatically map file columns to standard particle properties if names match
Bug fix: Particle selection in data inspector is lost when playing animation or moving viewport camera
Workaround for macOS (Apple Silicon) OpenGL stencil buffer issue: Highlighted particles not rendered correctly
Update third-party libraries: ffmpeg 6.0, OpenSSL 1.1.1u, libssh 0.10.5, Qt 6.5.1, PySide6 6.5.1.1, HDF5 1.14.1-2, NetCDF 4.9.2
Version 3.8.4 (03-May-2023)
Fix: ffmpeg video encoding crashes on Windows if output path contains non-ascii characters
Silence console message “Numeric mode unsupported in the posix collation implementation” on Linux by enabling ICU support in Qt build
pro Fix: Segfault in PySide6 package initialization on Linux when adding a Python layer to a viewport
py Fix: Interchanged xz/yz simulation box shear components in
lammps_to_ovito()Python function
Version 3.8.3 (16-Apr-2023)
Further improved performance of sequential loading of compressed trajectory files
Fixed regression (since v3.8.0):
Viewport.render_anim()renders only first animation framepy Python exceptions raised in user-defined modifier functions are now propagated up the call chain to where the pipeline evaluation was triggered
pro Included
bz2and sqlite3 standard modules, which were missing in embedded Python interpreter on Linux
Version 3.8.2 (04-Apr-2023)
Implemented fast access to trajectory frames in compressed (gzipped) files
Fix: Segfault when using zoom function in viewport with an attached camera object
Fix: Segfault in Coordination polyhedra modifier on Linux
Fix: Function ‘load/save session state’ does not follow global working directory
Version 3.8.1 (27-Mar-2023)
Identification of volumetric regions using the Gaussian density method pro
The Construct surface mesh modifier’s implementation of the Gaussian density method has been extended to support the identification of volumetric regions, e.g. pores, cavities, and filled spatial regions. Their respective surface areas and volumes are calculated and output by the modifier in tabulated form.
To make this possible, we have developed an extension to the Marching Cubes algorithm for isosurface construction, which provides the capability to identify disconnected spatial regions separated by the surface mesh and compute their enclosed volumes – of course with full support for periodic boundary conditions.
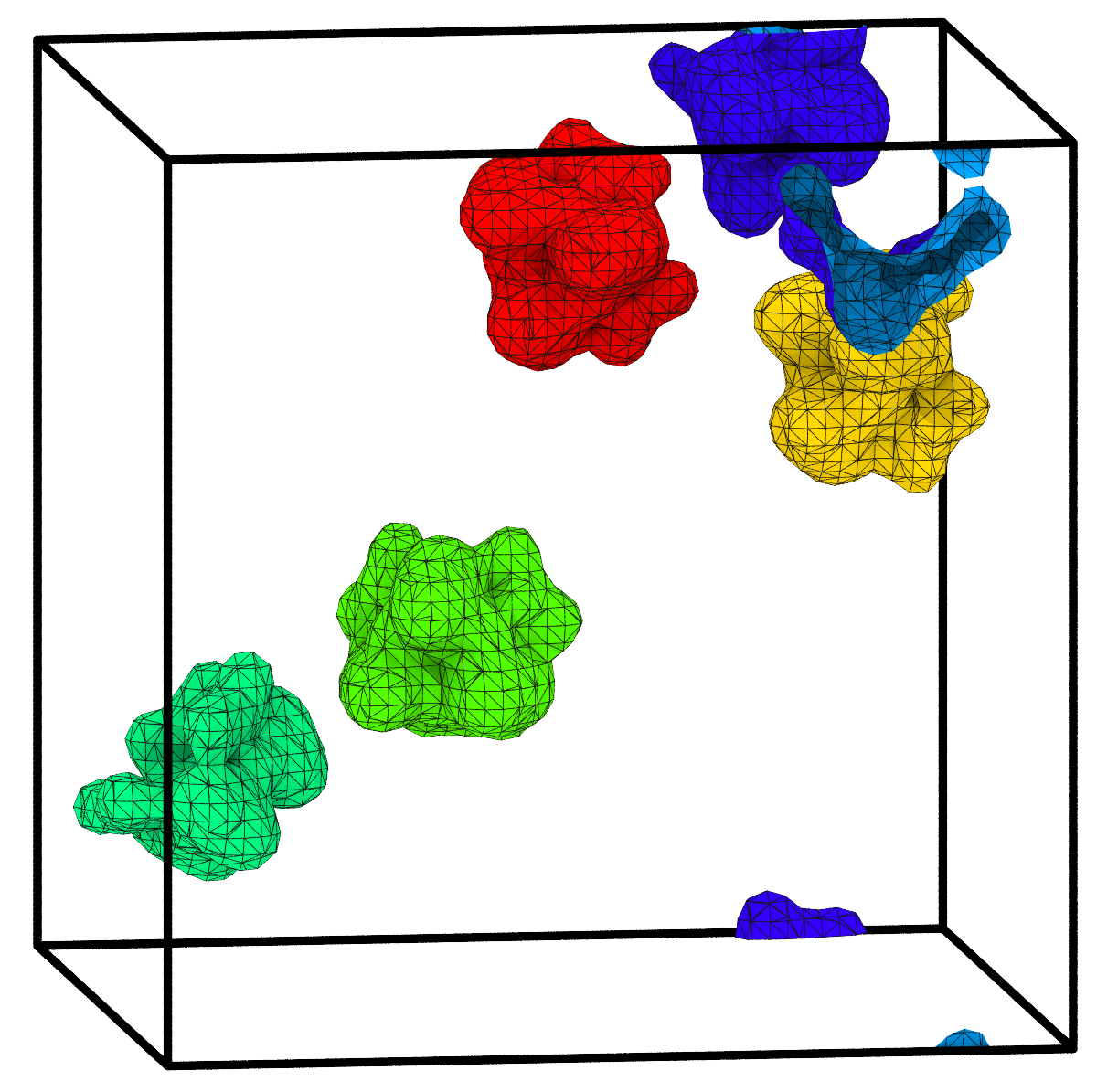
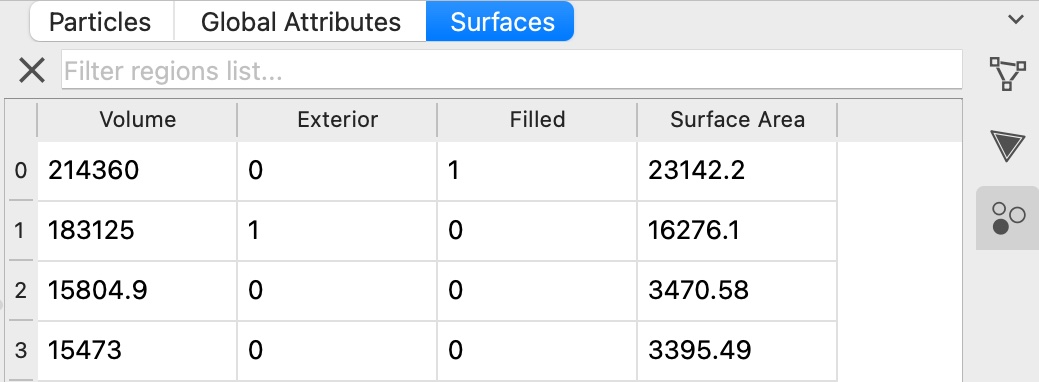
New efficient Python method for computing neighbor lists pro
OVITO’s Python interface now offers the new CutoffNeighborFinder.find_all() method
for vectorized computation of neighbor lists for many or all particles at once.
Further changes:
LAMMPS data file reader: Accept ‘#’ in type names, which are referenced in data sections of the file
Version 3.8.0 (03-Mar-2023)
Develop custom modifiers with extended capabilities pro
A newly devised programming interface enables you to write advanced modifier functions in Python that
access more than one frame of a simulation trajectory,
perform computations that involve data from several input files, or
need control over the caching of computational results.
Take simulation post-processing to the next level! Develop your own trajectory analysis algorithms in Python, which are fully integrated into OVITO’s pipeline system and the interactive interface of OVITO Pro.
class CalculateIncrementalDisplacementsModifier(ModifierInterface):
def modify(self, data, frame, input_slots, **kwargs):
next_frame = input_slots['upstream'].compute(frame + 1)
displacements = next_frame.particles.positions - data.particles.positions
Have a look at our completely revised introduction to user-defined modifiers and check out the new advanced programming interface for user-defined modifiers.
Improved color legends
OVITO can now render tick marks in color mapping legends to label intermediate values. Furthermore, the legend’s title may be rotated by 90 degrees:
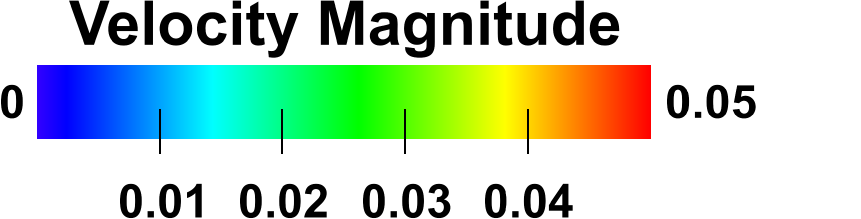
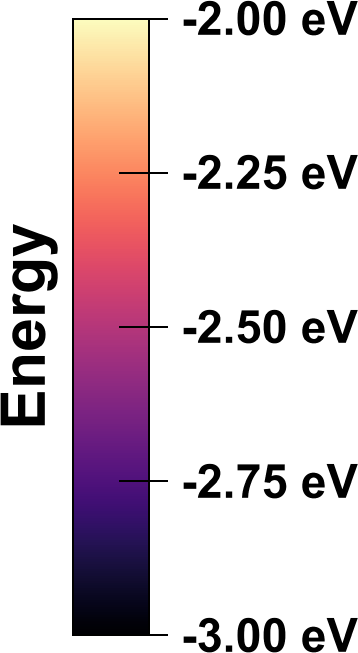
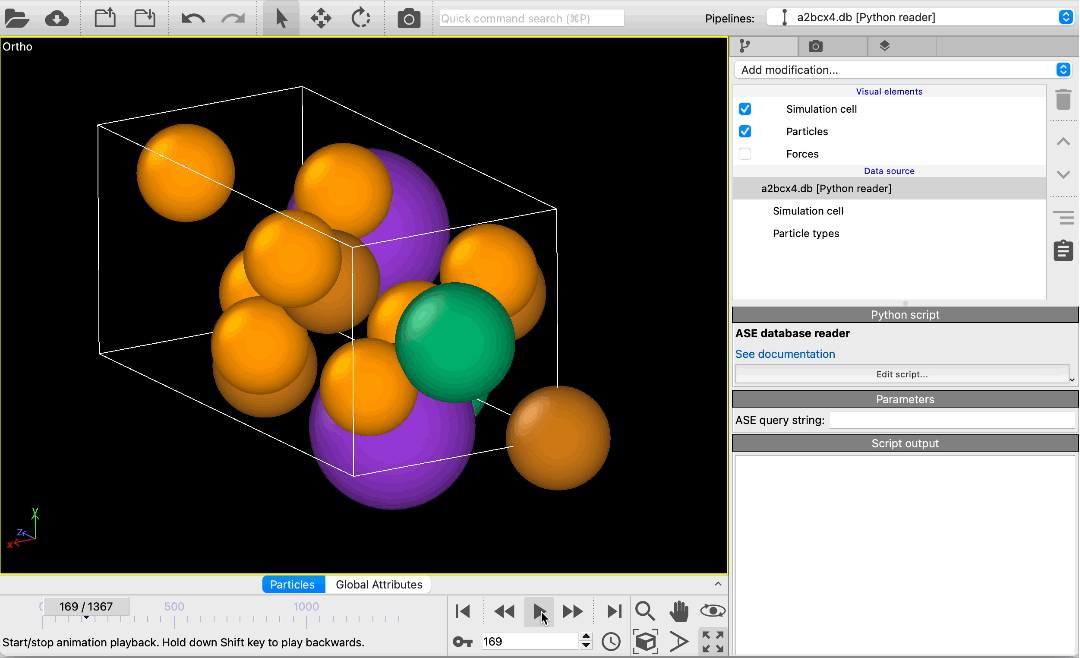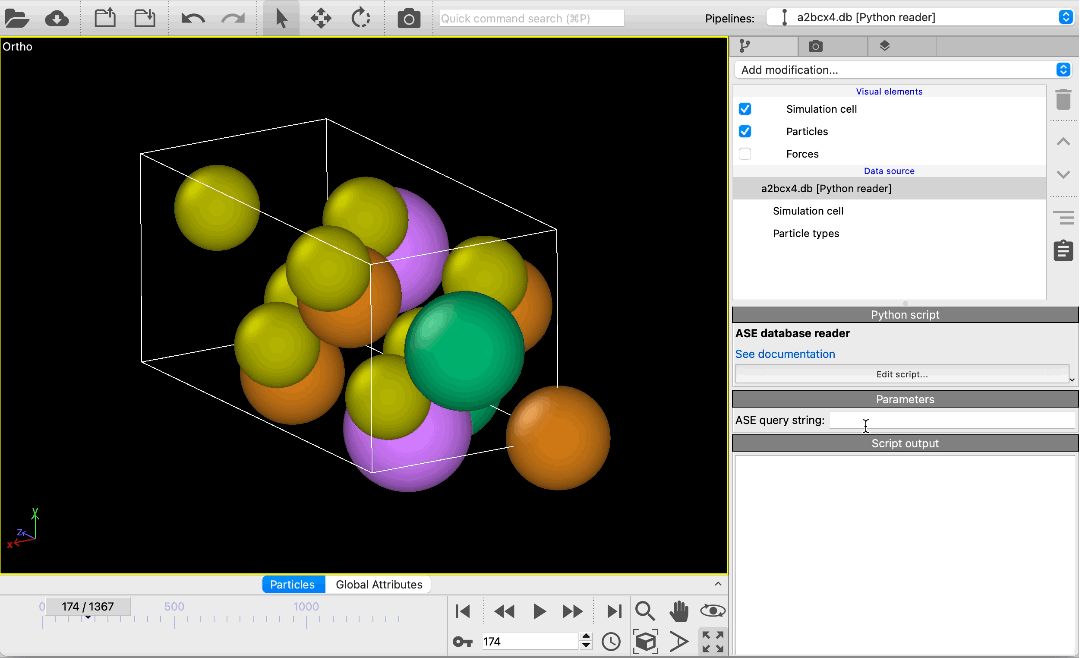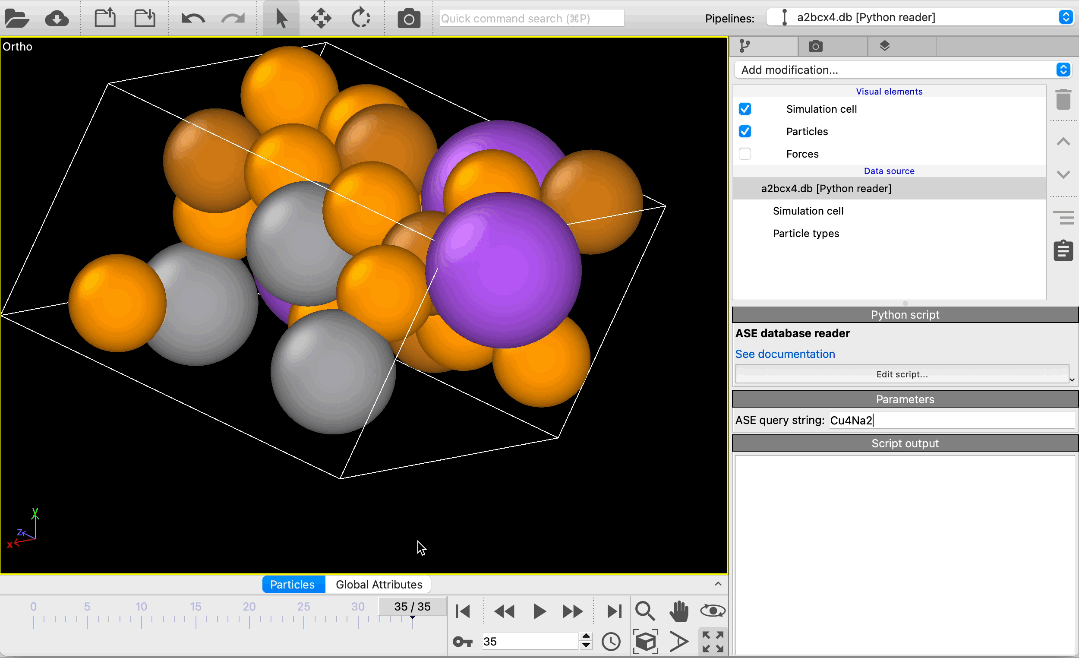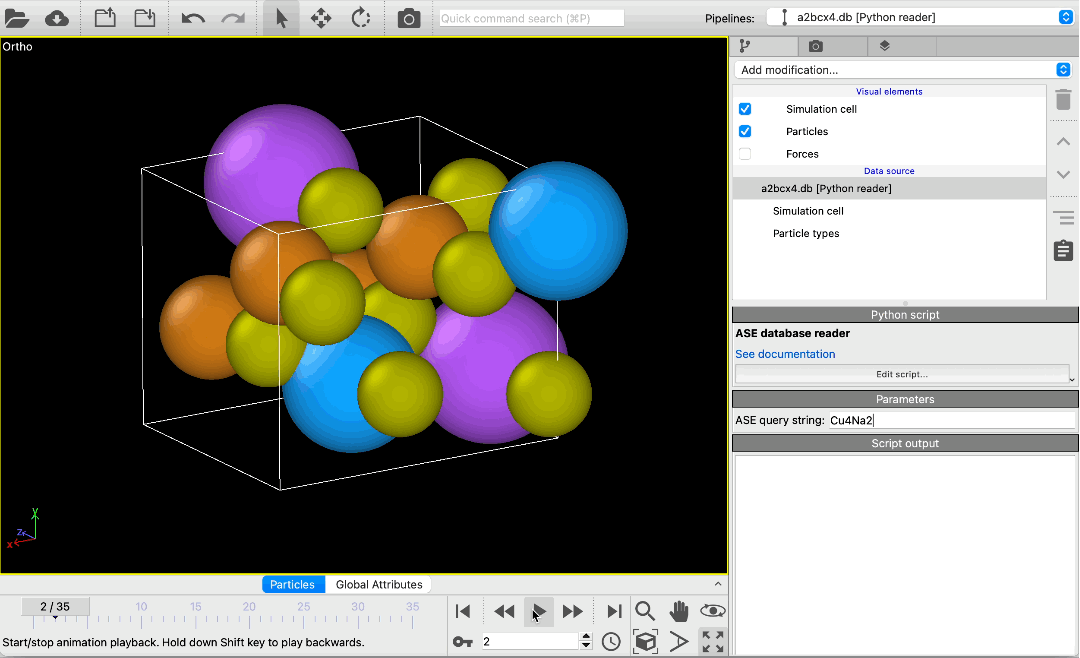
File reader for ASE database files pro
Load atomic structures from database files of the Atomic Simulation Environment (ASE) into OVITO. The new file reader lets you scroll through all structures in a database or pick specific structures using a query string. Metadata associated with structures is made available in OVITO as global attributes.
New modifier: Identify fcc planar faults pro
Easily identify different planar defect types, such as stacking faults and coherent twin boundaries, in face-centered cubic (fcc) crystals. We have developed a powerful classification algorithm for hcp-like atoms that make up such planar defects:
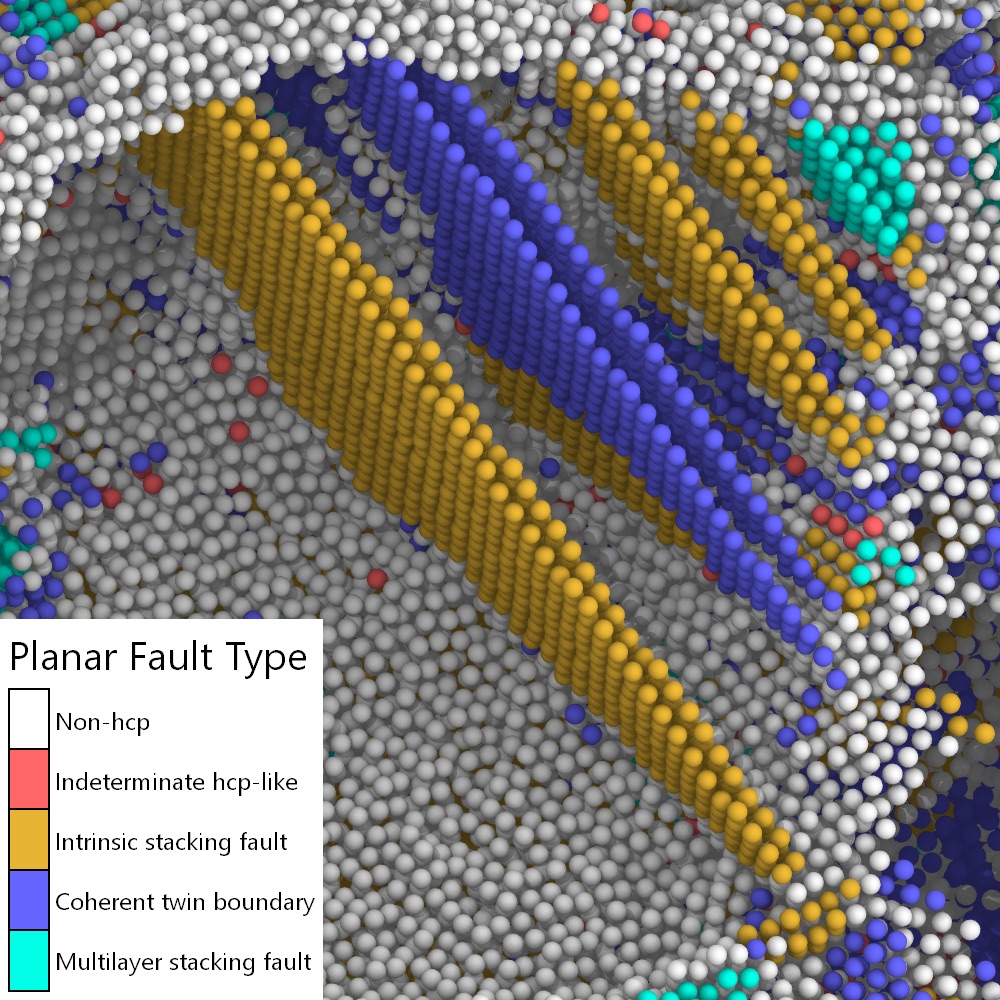
New modifier: Render LAMMPS regions pro
Use this new tool to generate mesh-based representations of the parametric regions defined in your LAMMPS simulation, e.g., cylinders, spheres, or blocks, and visualize the boundaries of these spatial regions along with the particle model:
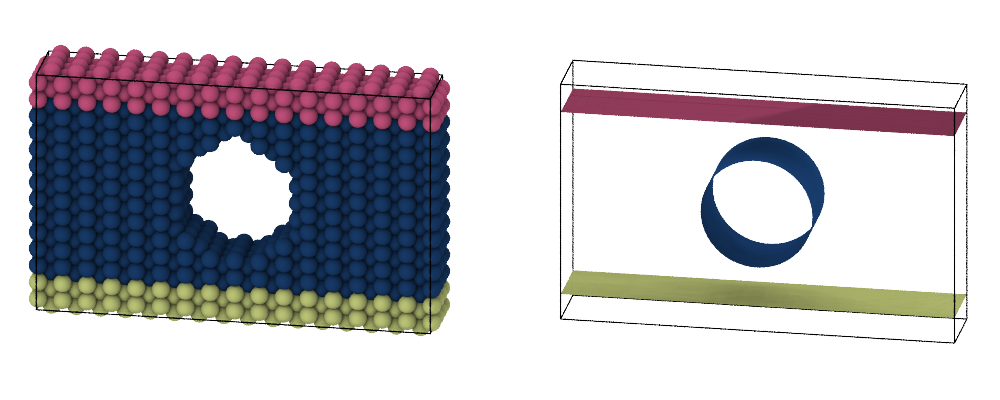
Spatial binning modifier: New unity input option pro
This options offers a shortcut for calculating particle density distributions, i.e. counting the particles per grid cell. Previous versions required first defining an auxiliary particle property with a uniform value of 1 to calculate the number density:
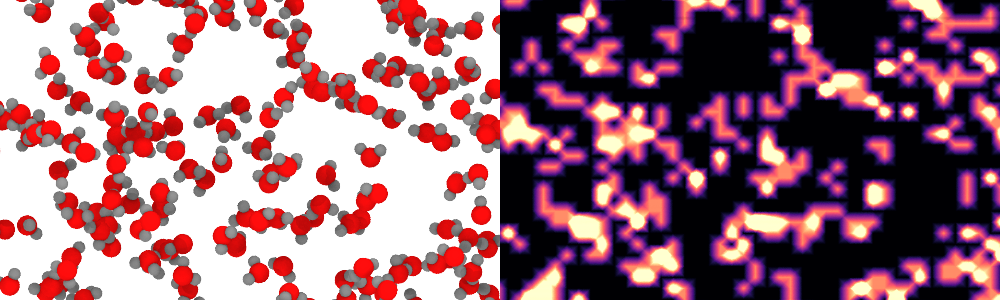
See Spatial binning modifier.
Support for LAMMPS dump grid files
OVITO can now read and visualize the new volumetric grid file format written by recent LAMMPS versions thanks to the newly added LAMMPS dump grid file reader:
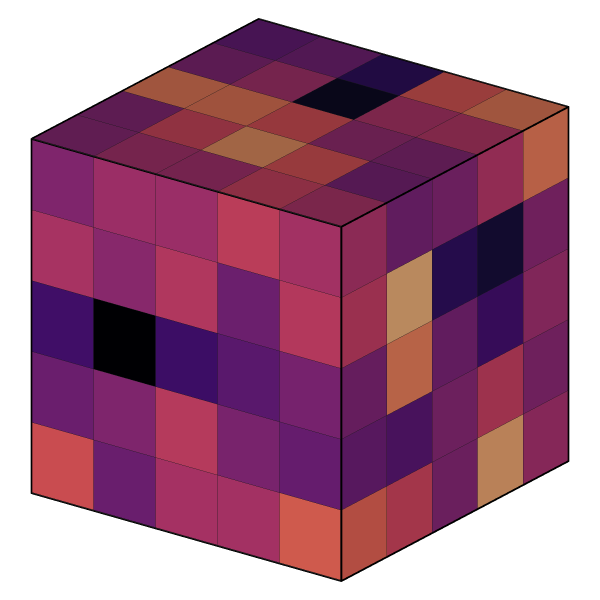

Slice modifier on voxel grids
When you apply the Slice modifier to a voxel grid, cell values now get copied to the mesh faces and interpolated field values to the mesh vertices of the generated cross-section. This enables both discrete and interpolated visualizations of the field values along arbitrary planar cross-sections:
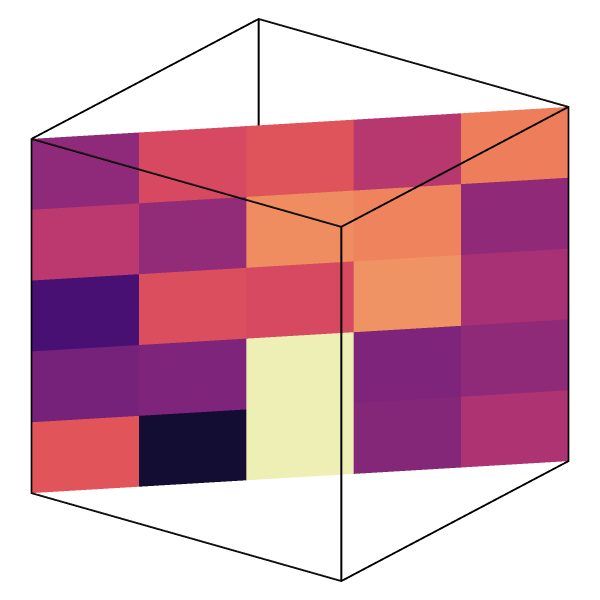
See Slice modifier and Voxel grids.
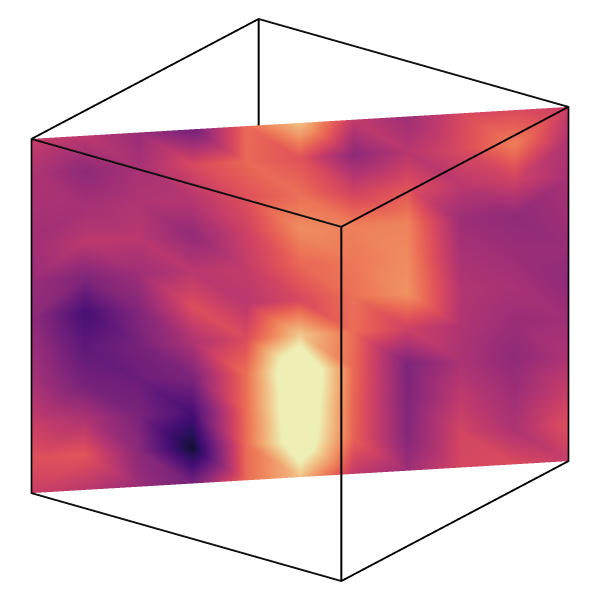
Support for point-based volumetric grids
In addition to the classical cell-based voxel grids, OVITO now also supports point-based volumetric grids, in which field values are associated with the grid points instead of the voxel cells. All functions in OVITO that operate on grids, e.g. the Create isosurface modifier, also support periodic and mixed boundary conditions.
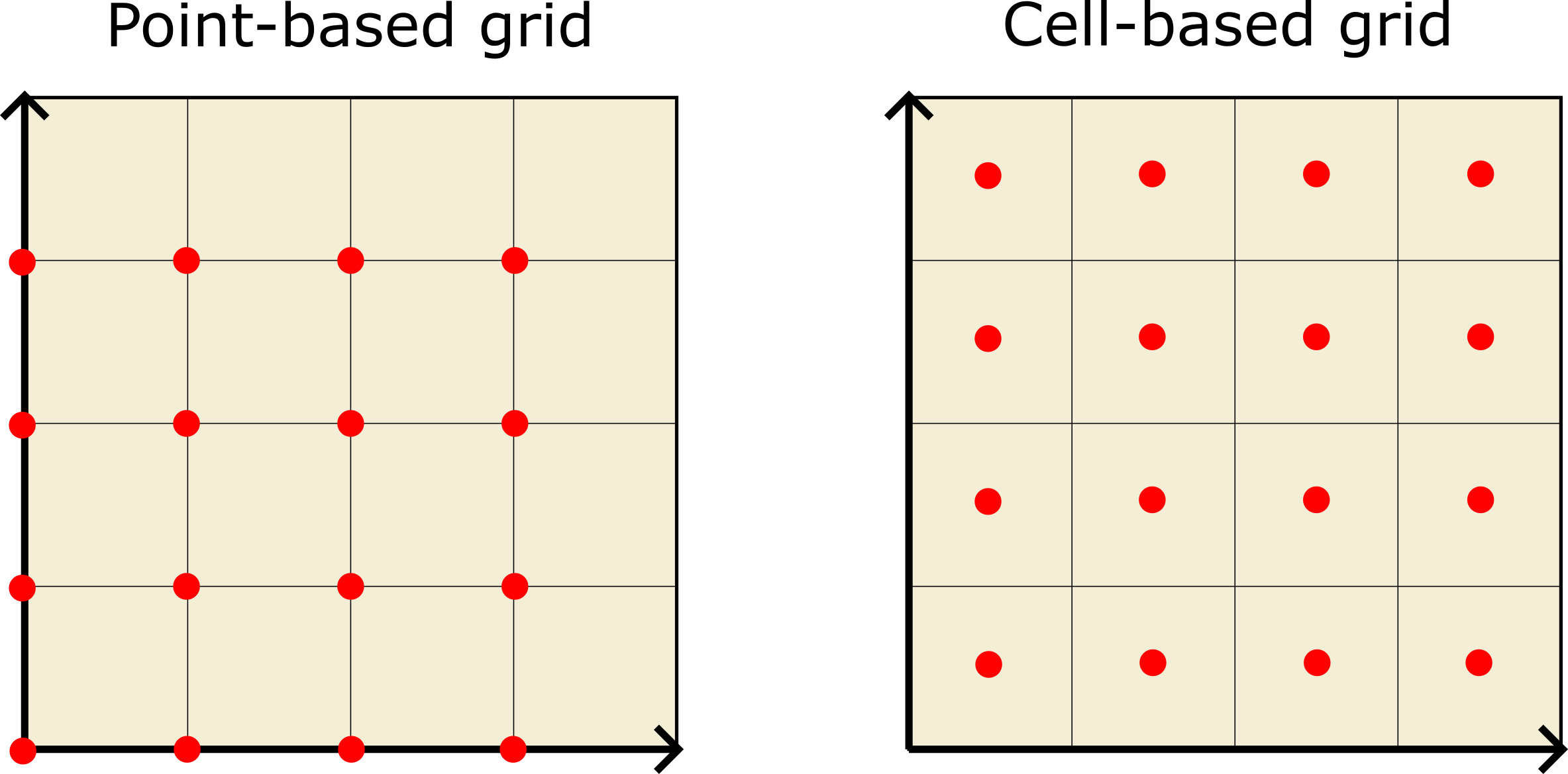
See ovito.data.VoxelGrid.grid_type and Gaussian Cube file reader.
Load Trajectory modifier now supports removal of particles
Previously, the Load trajectory modifier required the trajectory file to contain coordinates for all particles that were initially present in the topology dataset. The improved version of the modifier can now deal with particles disappearing in later frames of a trajectory, e.g., when particles get removed from the simulation over time.
Further additions and changes in this program release:
Added dark mode UI support for Linux platform.
Spatial correlation function modifier: Added support for 2d simulations.
Wrap at periodic boundaries modifier: Added support for 2d simulations.
Save and restore maximized state of main window across program sessions.
LAMMPS data file reader & writer: Added support for extended Velocities file section for when using LAMMPS atom styles electron, ellipsoid, or sphere.
LAMMPS data file writer: Added the option to renumber all particle/bond/angle/dihedral/improper types during export. Avoids conversion problems from 0-based type IDs loaded from GSD files.
New option to clip surfaces at open box boundaries (see
SurfaceMeshVis.clip_at_domain_boundaries).Cluster analysis modifier: Abort calculation of center of mass and radius of gyration if masses of all input particles are zero.
pro Added user option that makes OVITO Pro import multiple files of the same kind as separate objects into the scene.
py Accept
os.PathLikeobjects in Python functionsimport_file()andexport_file().py
PropertyContainer.create_property: Acceptdatavalues that are broadcastable to shape of property array.
Version 3.7.12 (16-Dec-2022)
GRO file reader: Recognize additional chemical symbols SI, FE, BR.
STL file reader: Tolerate leading whitespace on first line.
Updated third-party libraries on Windows: Qt 6.4.1, OpenSSL 1.1.1s, ffmpeg 4.2.8, zlib 1.2.23.
Fix: Voronoi cavity radius calculation is wrong by a factor of 2.
Fix: Function “Make Independent” does not work correctly for surface mesh visual elements in cloned pipelines.
pro Fix: Python method
ovito.data.SurfaceMesh.locate_point()can yield wrong results for coarse, one-sided meshes.
Version 3.7.11 (29-Oct-2022)
Added user option to application settings dialog for changing the working directory behavior.
Fixed regression: Slice modifier does not work on voxel grids.
pro Vectorized all query methods of
SurfaceMeshTopologyclass.pro Provide PyPI package for Python 3.11.
pro Added flat array option to method
SurfaceMesh.get_face_vertices().
Version 3.7.10 (09-Oct-2022)
Optimization of main window UI widgets to improve rapid animation playback at high frame rates.
Enhancements to the pipeline editor: Brief information display for some modifiers.
New right-click context menu in pipeline editor: Branched and cloned pipelines Added ‘Copy to pipeline…’ function for copying modifiers within and across pipelines.
pro Standalone Python module: Run in headless mode by default. OVITO_GUI_MODE env variable requests OpenGL renderer support.
pro PyPI package on Linux: Switched back to PySide6 version 6.2.4 for better backward compatibility with older Ubuntu distros.
pro Fixed loading of files opened via double click in case license validation dialog pops up.
pro Generalized the Vectors element to support visualization of vector quantities in more types of
PropertyContainers.pro New Python function
ovito.modifiers.PolyhedralTemplateMatchingModifier.calculate_misorientation().pro Automatic conversion of NumPy array scalars to Python numbers when storing them as OVITO global
ovito.data.DataCollectionattributes.
Version 3.7.9 (12-Sep-2022)
Voronoi analysis: Added calculation of cavity radius.
GSD file importer/exporter: Added support for particle attributes “angmom” and “body”.
Fix: Affine Transformation modifier not transforming particles in target-cell mode in rare situations (when called from Python).
Fix: File import via drag & drop not working when Vulkan viewport renderer is active.
Upgraded Qt cross-platform framework to version 6.3.1.
Cluster analysis modifier: Warn user if center of mass cannot be computed due to cluster’s total mass being zero.
pro Upgraded OVITO Pro integrated interpreter embedded interpreter to Python 3.10.6.
pro Installing third-party Python modules Installation of PyPI packages with the
--useroption in the embedded interpreter is now supported.pro New Python API for creating
ovito.data.SurfaceMeshobjects.pro Improved operation of Python module in Jupyter environments. Interrupting long-running operations is fully supported now.
pro New experimental Jupyter notebook visualization widget (
ovito.vis.Viewport.create_jupyter_widget()).pro Added Python API
ovito.vis.ColorLegendOverlaycolor_mapping_source.pro Fix: Segfault during Python statement
del ovito.scene.pipelines[:].
Version 3.7.8 (29-Jul-2022)
Fix: Program crash when quickly skipping through a trajectory consisting of a series of files loaded via SSH (regression OVITO 3.7.0).
Fix: Visual artifacts when rendering cone primitives (3d arrow heads) at small length scales due to numerical precision issue.
pro Added conda packages for Python 3.10.
pro Added conda packages for macOS arm64/M1 platform.
pro Work around a memory leak in some OpenGL graphics driver implementations when the
ovito.vis.Viewport.render_image()Python function is called repeatedly.
Version 3.7.7 (06-Jul-2022)
Ubuntu 22.04 compatibility - Linux package of OVITO now includes a private copy of OpenSSL 1.1 libraries.
Version 3.7.6 (23-Jun-2022)
PDB file reader: Added support for CP2K trajectory format.
LAMMPS dump file reader: Recognize
quat{ijkw}andshape{xyz}columns and automatically them to correct particle properties.Fix: Camera FOV parameter not animatable when rendering a movie.
Fix: Segfault when loading .ovito state files written by OVITO 3.3 or older containing a Python script.
Fix: Grain segmentation algorithm never terminates for particular inputs.
PyPI package for Linux: disabled built-in SSH client to improve compatibility with Ubuntu 22.04, which doesn’t provide OpenSSL 1.1 libraries anymore.
pro New Python class
ovito.data.SurfaceMeshTopology, which provides script access to the face connectivity information of surface meshes.pro Conda channel now provides additional variants of the ovito package (built against
tbbv2020 and v2021), which avoids dependency conflicts with certain third-party packages when installing them in the same environment.
Version 3.7.5 (28-May-2022)
Smooth trajectory modifier now supports varying number of particles.
SSH client: Try password first before keyboard-interactive authentication for successful handshaking with some SSH servers.
Performance improvements to OpenGL high-quality sphere rendering code
Bug fix: Data inspector shows a 3rd text label in bar charts with 2 bars.
Bug fix: Sporadic program crashes when importing CA files.
pro
DataCollection.attributesdictionary can now store arbitrary Python objects.pro New Python method
ovito.data.Particles.remap_indices().pro New Python method
ovito.data.SurfaceMesh.to_triangle_mesh().pro Bumped maximum neighbor limit of
ovito.data.NearestNeighborFinderto 64.pro Dropped support for Python 3.6, which has reached its end-of-life date.
Version 3.7.4 (18-Apr-2022)
Centrosymmetry parameter modifier: New option ‘Use only selected particles’.
LAMMPS data file reader: Added support for Ellipsoids section.
Fix: Program crash during file format detection when importing file from path containing CJK or other non-ANSI characters.
Fix: Error “The file source path is empty or has not been set” when picking a new simulation file of different format.
pro Construct surface mesh modifer: New option ‘Map particles to regions’.
pro New Python methods
ovito.data.DataCollection.create_cell(),ovito.data.DataCollection.create_particles(),ovito.data.Particles.create_bonds().
Version 3.7.3 (29-Mar-2022)
DXA modifier now picks up partitioning established by Grain Segmentation modifier in the upstream pipeline, see discussion in the forum.
Fix: XYZ file column mapping is reset when using “Pick new file” function.
Fix: App closes when using the “Pick new file” function under Linux (issue #216).
Fix: Segfault when deleting a disabled modifier from a branched pipeline.
Fix: Construct surface mesh modifier sometimes produces incorrect cap polygons if the alpha-shape complex contains degenerate elements (issue #217).
Regression: Progress bar not updated correctly during execution of Construct Surface Mesh and DXA modifiers.
Regression: Program does not exit if
--helpcommand line option is used.pro Added user documentation for Python-based modifiers Calculate local entropy and Shrink-wrap simulation box .
pro Added .pyi stub files to OVITO Python Reference Python package to support auto-completions and mouse-over documentation in Python IDEs.
pro
ovito.data.CutoffNeighborFindernow accepts non-periodic simulation cells that are degenerate.pro Fix:
ovito.data.DataTable.xy()method generates wrong x-coords array if data table interval doesn’t start at 0.
Version 3.7.2 (03-Mar-2022)
Improved render output window with image zoom function.
Fix: Particle type colors not initialised correctly if imported LAMMPS dump file contains both ‘type’ and ‘element’ columns (issue #193 note 403792737).
pro macOS: Fixed PySide6 loading error due to wrong rpath information when importing PyPI ovito package.
pro Linux: Fixed sqlite3 Python package included in the embedded Python interpreter of OVITO Pro.
Version 3.7.1 (26-Feb-2022)
Fixed regression: Segfault when loading session state file containing a viewport camera object.
pro New Python function
ovito.data.NearestNeighborFinder.find_all().pro
ovito.data.PropertyContainerclasses support removing properties with thedelstatement.pro Inform user if insufficient file access permissions let license activation fail.
Version 3.7.0 (15-Feb-2022)
Visual element and particle type settings can now be preserved when picking a new input simulation file in the External file external file panel.
Support for HTML formatted text in viewport layers Text label layer, Color legend layer, and Coordinate tripod layer.
Improved color quality of animated GIFs produced by OVITO
Added dark mode UI support on macOS.
Availability of native arm64/M1 builds of OVITO Basic, OVITO Pro and the OVITO Python package for Apple Silicon machines.
Ported OVITO code base from C++14 to C++17 language standard.
Switched from old Qt 5.x to version 6.2 of the Qt cross-platform C++ framework and, correspondingly, from PySide2 to PySide6. (Exception: Packages for Anaconda, where dependencies Qt6/PySide6 are not yet available).
Completely reworked and modernized the internal asynchronous task system and the scene rendering framework of OVITO.
New standard Bonds property “Width”, which allows controlling the diameter of bond cylinders on a per-bond basis.
Added detailed documentation for some of the file readers of OVITO. See the Input file formats.
LAMMPS data file reader & writer: Added preliminary support for type labels, which will be supported by a future version of LAMMPS.
LAMMPS dump file reader: Map columns
c_diameter[...]to particle propertyAspherical Shapeand perform division by 2.GSD file reader & writer: Added support for angles/dihedrals/impropers.
User can now rename individual structure types in the UI of structure identification modifiers.
Implemented new OpenGL rendering technique *Weighted Blended Order-Independent Transparency*, providing an alternative to the classical painter’s algorithm. Can be activated in the app settings dialog and gives better results if there’s a mix of several different object types (e.g. particles and surfaces) that are all semi-transparent.
Detect if the triangle mesh is not closed when loading a custom particle shape. Automatically disable back-face culling for the particle type in this case.
CA file reader: Compute dislocation line statistics for re-imported datasets the same way the DXA modifier does.
Fix: Particles visual element does not use uniform scaling factor when rendering some non-spherical particle shapes.
Version 3.6.0 (19-Nov-2021)
Vectors, Surface mesh, Voxel grid, Lines visual elements: Added direct color mapping option as a faster alternative to the Color coding modifier.
Bonds visual element: Added explicit control of the coloring mode.
Made number and Multi-viewport layouts configurable by the user.
Visibility of pipelines can be controlled on a Viewport menu per viewport basis.
Coordination polyhedra modifier now makes particle properties available for the color coding as mesh region properties and mesh vertex properties.
Generate trajectory lines modifier: New capability to transfer time-dependent particle properties to the trajectory lines.
Load trajectory modifier: Support non-contiguous atom IDs in LAMMPS bond dump files
Added file reader for binary STL files.
Customizing the initial program state: New mechanism for customizing the initial program session state.
Raised limit on the number of FFT bins in Spatial Correlation Modifier to support finer grid resolutions.
Fix: PTM modifier may crash if graphene/diamond are the only enabled structure types.
Fix: Traced trajectory lines may be rendered in wrong colors.
Took out code that transmits random installation ID to web server.
OpenSSL shared libraries are no longer shipped with OVITO for Linux to avoid compatibility issues on some Linux distributions.
pro Rendering images or movies of a viewport layout : Added capability to render multi-viewport layouts in one step.
pro Python code generator has been extended to generate code for all visual elements and for reenacting manual changes made by the user to data objects (e.g. particle type names, color, radii).
pro Added the
input_formatkeyword parameter to theovito.io.import_file()Python function for specifying the file format explicitly.pro Upgraded OSPRay to version 2.7.1.
pro Renamed
ovito.vis.Viewport.create_qt_widget()method and made it work in all distributions of theovitoPython module.pro Added experimental
ovito.vis.Viewport.create_jupyter_widget()method for embedding OVITO viewports in Jupyter notebooks (see demo binder).pro Support for site-wide software licenses.
pro Fix: Bounding box clipping artifact when rendering rotated superquadrics particles with OSPRay or Tachyon renderers.
pro Fix: Warning “This plugin does not support createPlatformOpenGLContext!” when running in headless mode on Linux machines.
Version 3.5.4 (31-Jul-2021)
LAMMPS data file reader and writer now support all LAMMPS atom styles, including the
hybridstyle.Fix: Construct surface mesh with region identification fails or never completes for some inputs.
pro Fix: Tachyon renderer crashes when triangle mesh contains a degenerate vertex normal.
Version 3.5.3 (30-Jun-2021)
Added two Tutorials to the documentation.
Voxel grid visual element now supports mouse-over data display in the status bar.
Added invert function to Manual selection modifier.
Warn user if OVITO Python module was installed via
pipcommand in an Anaconda Python interpreter. Useconda installinstead!Fix: Configure Trajectory Playback dialog shows no contents.
Fix: Neighbor finder facilities do not ignore PBC flag along third dimension in 2D mode.
Version 3.5.2 (26-May-2021)
Affine transformation modifier now allows entering the translation vector in reduced cell coordinates.
Load trajectory modifier can now import ReaxFF bond information files written by the LAMMPS fix reax/c/bonds command.
GSD file reader: Fill particle property array with default values if a chunk is not present in current frame (issue #206)
pro Fix: Invisible simulation cell edges when rendering image with orthographic projection with OSPRay
Version 3.5.1 (18-May-2021)
The Radial distribution function (RDF) modifier has gained an option ‘Only selected particles’, which restricts RDF calculation to a subset of particles.
The ‘Generate neighbor bonds’ option of the Voronoi analysis modifier is now able to deal with small periodic simulation cells.
Fix: Wireframe line rendering issue in perspective viewports.
pro The Slice modifier now accepts (hkl) Miller indices as input for defining the plane orientation. The plane position can be specified in terms of the interplanar spacing.
pro OVITO Pro for Linux now ships with a current Python 3.9.5 interpreter.
pro Fix:
ovito.data.PropertyContainer.create_property()method cannot create user-defined property of data typeint64.
Version 3.5.0 (02-May-2021)
Pipeline editor supports drag-and-drop operations, which allow easy rearranging of modifiers with the mouse.
Modifiers can be grouped in the pipeline editor to collapse complex sequences of modifiers into a single list entry.
Color legend layer can render a legend for typed particle properties, showing the discrete colors representing the defined particle types.
New implementation of the OpenGL viewport renderer. Provides better compatibility with GPU hardware, older OpenGL drivers, and virtual machine environments. OVITO now works on systems with only OpenGL 2.1 support (previous OVITO version required OpenGL 3.2).
New viewport renderer based on the Vulkan graphic hardware interface as an alternative option to the OpenGL renderer. Can be activated in the application settings dialog (not available on macOS). Supports rendering in head-less mode on HPC nodes with GPU hardware.
New Pipeline concept pipeline selector widget in the toolbar of OVITO, which lets you manage the data pipelines in the current scene and add new pipelines.
Extended the Create bonds modifier. A new parameter-free mode allows creating bonds based on van der Waals radii of the atoms.
Performance improvement: Create Bonds modifier can now make use of multiple processor cores.
Affine Transformation modifier can now transform triangle meshes (imported from STL, OBJ, VTK files).
Several file format readers now provide the option to generate interatomic bonds during data import (relieves from having to apply the Create Bonds modifier).
Some file format readers provide a new option to dynamically recenter the simulation cell on the coordinate origin. Useful for visualizing trajectories with varying cell shape.
Gromacs, PDB, and mmCIF file readers now import atom names and residue names as particle properties.
Internal chemical database of OVITO has been extended to include all elements and mass information, which will be assigned to particle types during file import.
The Particles visual element provides a Particles new parameter controlling the uniform scaling of atom radii. Useful for quickly producing a typical “balls-and-sticks” representation of a molecular structure.
A bonds-only visualization of a molecular structure (with particles turned off) now adds spheres at the nodal points of the bond network to yield a typical “stick” representation.
XYZ file reader now supports the exyz format variant of OpenBabel.
Fix: CFG file reader loosing particle type settings during file reload.
Fix: Segfault when loading certain NetCDF files with >1M particles.
Fix: Error when deleting some regions of a surface mesh structure.
Fix: Slow performance of Particles visual element when some particle types use mesh-based shapes.
Rearranged the Render settings panel. The viewport preview mode can now be activated from here.
Extended the Python API to support
ovito.data.VoxelGridfrom scripts.OVITO User Manual uses a new layout theme and supports full-text search.
Environment variable OVITO_LOG_FILE allows redirecting terminal output of OVITO to a text file (useful on Windows platform, where console output is otherwise inaccessible).
pro New modifier Color by type modifier for recoloring particles based on one of their typed properties, e.g. discrete
Residue TypeorAtom Nameproperty.pro New pipeline data source type Python script . Run a user-defined Python function that builds or synthesizes an input
DataCollectionfor a pipeline (instead of loading a structure from disk). Can also be used to import data formats into OVITO which are not directly supported by the software.pro LAMMPS script : The new data pipeline source type LAMMPS script allows editing and executing LAMMPS input scripts within OVITO to generate a dataset using LAMMPS commands. Useful for prototyping LAMMPS simulation setups with immediate visual feedback in OVITO.
pro Updated OSPRay rendering library to version 2.5.0, offering a better denoising filter.
pro Spatial Binning modifier can now process vector particle properties in addition to scalar properties.
pro Python API: Added the method
ovito.data.CutoffNeighborFinder.find_at()for enumerating all particles around an arbitrary spatial position.pro Python code generator: Emit valid code for visualization setups including a
PythonViewportLayer.pro Python code generator: Emit call to
generate()method of Generate Trajectory Lines modifier.pro Fix: Made auto-crop function work for pictures rendered with OSPRay and denoising filter enabled.
pro Fix: Python viewport layer does not get called with current values of user-defined parameters.
Version 3.4.4 (12-Mar-2021)
Fix: Number of data columns not correctly detected for XYZ files with 5 atoms or less.
Fix: Program crash when playing back animation with less than 1 frame per second in interactive viewports.
Fix: Simulation cell not visible in interactive viewports on some computer systems (issue #203).
Fix: CIF file reader not automatically recognizing files written by Open Babel (issue #204).
pro Fix: OSPRay not rendering arrow glyphs correctly.
Version 3.4.3 (25-Feb-2021)
Added text outline option to Coordinate Tripod viewport layer.
Fixed UI issue: Status bar resizing due to invalid unicode character in text string.
Corrected camera orientation of “Bottom” viewport view type when rotation constraint is turned on.
Improved automatic detection of PDB file format.
pro It’s now okay to assign a simple string to the
ovito.modifiers.ExpressionSelectionModifierexpression field.
Version 3.4.2 (15-Feb-2021)
Long text strings displayed in the status bar of OVITO now get broken into two lines in order to show more property values in the available space.
Bug fix: Status bar doesn’t display latest set of particle properties while positioning the mouse cursor over a particle. This fix corrects a regression introduced with OVITO 3.4.0.
Fixed a limitation of the PTM modifier not identifying diamond and graphene structures in small periodic simulation cells.
Version 3.4.1 (03-Feb-2021)
Fixed runtime linker error when importing
ovitoPython module installed via pip on Linux.pro Spatial binning modifier can now operate on vectorial particle properties.
Version 3.4.0 (28-Jan-2021)
Backward incompatible .ovito state file format change: Program sessions saved with OVITO 3.4 or later cannot be opened in previous versions!
Extensive redesign of OVITO’s internal C++ data object model to make it thread-safe. User experience and Python API remain largely unaffected.
State files (.ovito) now store relative paths to imported data files, enabling the relocation of an entire directory tree containing the state file and the data file without breaking the reference.
Rewrite of the OpenGL rendering code, making use of geometry shaders on a wider range of hardware. OVITO now requires OpenGL 3.0 or higher (previous releases required OpenGL 2.1).
Color Coding modifier: Added an auto-adjust option, which dynamically adjusts the min/max interval to the current range of input values.
File importers reading the
Velocityvector particle property automatically generate theVelocity Magnitudeparticle property too.OVITO can now visualize particles with superellipsoid shapes, which are controlled by the
Superquadric Roundnessparticle property.Preliminary file reader support for ParaView VTP, VTI, VTM and PVD formats, as written by the Aspherix DEM simulation code.
pro OVITO Pro gives the user the option to edit Python scripts in an external editor application or IDE (e.g. Visual Studio Code). Changes the user makes to the script code in the external editor are automatically loaded back into OVITO Pro.
pro The Python script modifier displays the current working directory and lets the user control it if necessary.
pro Python-based viewport layers now support user-defined parameters passed to the
render()function.pro OSPRay and Tachyon renderers can now render polyhedral meshes with highlighted edges (wireframe overlay).
pro New Python method
ovito.data.NearestNeighborFinder.find_at().
Version 3.3.5 (12-Dec-2020)
Extended the Smooth Trajectory modifier to interpolate/average all scalar and continuous particle properties.
Fixed handling of stacking faults of arbitrary thickness in grain segmentation algorithm.
pro Fixed shading issue for ellipsoidal particles in OSPRay renderer.
pro Fixed z-clipping issue in OSPRay and Tachyon renderers for viewports with parallel projections.
Version 3.3.4 (27-Nov-2020)
Another tweak to the PTM algorithm to fix a regression, which let the PTM modifier fail to correctly identify some BCC atoms.
Version 3.3.3 (23-Nov-2020)
Construct Surface Mesh modifier: New capability to compute distance of each particle from closest point on the surface.
Added a user option to Text Label viewport layer, which allows controlling the output precision and formatting of decimal values.
Added support for the improved binary dump file format introduced with LAMMPS stable release 29-Oct-2020.
Fixed an issue in the PTM algorithm letting the identification of BCC atoms sporadically fail if exactly arranged on a perfect lattice.
pro Fixed visual issue in OSPRay renderer when rendering semi-transparent ellipsoidal particles.
pro New code example showing how to access partial RDFs computed by CoordinationAnalysisModifier.
Version 3.3.2 (12-Nov-2020)
Included shared library libxcb-xinerama.so in binary package for Linux, which may not be present on some systems by default.
Version 3.3.1 (13-Oct-2020)
Grain segmentation modifier: Fixed a bug in the handling of stacking faults.
pro Support license option that allows running the GUI on arbitrary nodes of a computing cluster.
Version 3.3.0 (07-Oct-2020)
New option in application settings dialog: Sort list of available modifiers by name instead of category.
Data plot window now support mouse interaction (zooming/panning), allowing you to take a closer look at the displayed graph.
Create isosurface modifier: Iso-value can now be set with the mouse by clicking into the histogram plot.
Create isosurface modifier: New option Transfer field values to surface for mapping the all voxel grid properties to the isosurface, which can subsequently be used to locally color the generated surface according to some secondary field quantity.
Slice modifier: Now supports slicing of voxel grids to extract planar cross-sections.
Added a quick search field for quickly accessing modifiers and other program commands in the GUI.
The file selection dialog now allows importing multiple files at once. In particular, combinations of topology and trajectory files can now be opened in one step, with OVITO inserting the Load Trajectory modifier automatically.
Added a file reader for the Gromacs GRO and XTC file formats.
Added option Ignore particle identifiers to LAMMPS data file exporter, which will reassign a contiguous range of identifiers to the exported atoms.
New implementation of the PDB file reader based on the third-party Gemmi library, providing better compatibility with a wide range of files.
Expression selection modifier: Expressions for selecting bonds can now reference properties of the two connected particles.
Viewport camera can now be controlled using arrow keys. Use shift key for panning.
OVITO is now based on Qt 5.15.1, fixing some UI issues on high-DPI screens under Windows.
pro New modifier: Time series - a convenient tool for plotting the time evolution of global simulation attributes in OVITO.
pro Time averaging modifier: Support multiple input quantities to be averaged simultaneously.
pro Spatial binning modifier: The output VoxelGrid object now has the identifier ‘binning’ instead of ‘binning[]’. Note that this represents a breaking change to existing Python scripts employing the SpatialBinningModifier class.
pro Bug fix: Python code generator includes ‘LAMMPSAtomStyle.’ prefix in
import_file()calls.
Version 3.2.1 (28-Aug-2020)
Load Trajectory modifier: Support trajectory datasets containing more particles than the topology dataset.
LAMMPS dump file reader: Support files containing both type and element data columns (issue #193).
OVITO for Windows: Fixed UI layout on high-DPI screens.
Upgraded third-party software components: Qt 5.15, Python 3.8.5, NetCDF 4.7.4, Libssh 0.9.4, OSPRay 2.2.0.
pro OSPRay: Applied patch to fix a memory leak (issue #196).
Version 3.2.0 (10-Aug-2020)
Added a file reader for the PDBx/mmCIF file format.
Vector visualization element: Added offset and transparency parameters.
Extended the Load Trajectory modifier to support loading dynamic bond topologies from dump local files written in LAMMPS reactive MD simulations.
New modifier: Interactive Molecular Dynamics (IMD) for live visualization of running MD simulations.
Cluster analysis modifier: Added calculation of weighted center of mass and radius of gyration.
LAMMPS dump file reader: Support files written with dump_modify time yes or dump_modify units yes.
pro New Bond Analysis modifier for calculating bond-angle and bond-length distributions.
pro New Python-based modifier for calculating the local entropy fingerprint proposed by P. M. Piaggi and M. Parrinello.
pro Python modifiers can now define their own function parameters. The values can be edited in the user interface and get stored in .ovito files.
pro Added the PropertyContainer.delete_elements() and delete_indices() methods for deleting particles, bonds, etc.
pro Added the CutoffNeighborFinder.neighbor_distances() and neighbor_vectors() Python methods.
pro Added the Viewport.create_widget() method for building simple user interfaces that make use of OVITO’s interactive visualization capabilities.
Version 3.1.3 (30-Jul-2020)
PTM modifier: Fixed identification of chemically ordered binary structures, which got broken in a recent update
PDB file format reader: Support for datasets with more than 9,999 atoms (see merge request)
Version 3.1.2 (13-Jul-2020)
New option for turning off automatic generation of file search patterns
New Configure Trajectory Playback dialog, allows controlling the mapping of trajectory frames to animation frames
GSD file reader: Now accepts ellipsoid shape definitions with principal axes b=0 and/or c=0
Bug fix: Animation rendering process cannot be canceled sometimes
pro Added DislocationNetwork.set_segment() method for manipulating dislocation line data from Python
Version 3.1.1 (21-Jun-2020)
Bug fix: LAMMPS data file reader fails to correctly read Masses file section with irregularly ordered atom types
pro Time averaging modifier: Check that x-values of data points are constant when averaging a data table
Version 3.1.0 (14-Jun-2020)
pro New feature: Python code generator
New feature: Grain segmentation modifier
Version 3.0.1 (05-Jun-2020)
Bug fix: Adjusted internal parameter of Construct Surface Mesh modifier to avoid sporadic error “Adjacent cell face not found” for periodic systems.
Enhancement: LAMMPS dump file reader can now parse “diameter” file column as Radius particle property, automatically performing division by 2.
Version 3.0.0 (30-May-2020)
Smooth Trajectory modifier interpolates particle orientations in addition to particle positions.
Added capability to Construct Surface Mesh modifier to identify pores/voids and individually compute their volumes and surface areas.
Extended the OSPRay renderer with a sky & sun light source.
Integrated copy protection into OVITO Pro builds.
Introduced OVITO_THREAD_COUNT environment variable for controlling the number of CPU cores used by OVITO.
Added old VASP 4.x file format to POSCAR reader (in order to support files written by ASE).
Added user option to LAMMPS data writer for omitting the “Masses” file section.
OVITO can now read/write extended topology information from/to LAMMPS data files: angles/dihedrals/impropers.
Bug fix: Output of Displacements/Atomic Strain/Wigner-Seitz modifiers can be all zeros in relative offset mode (dev679 regression).
Eliminated bottlenecks in GUI, which slowed down animation playback of long trajectories (>100k frames).
Correct handling of LAMMPS dump files with dual data columns (issue #193).
Updated OSPRay renderer to version 2.1.0 and integrated OSPRay plugin into Anaconda build of OVITO.
Bug fix: Color Coding modifier does not list available bond properties.
Added HTTP protocol support. OVITO can now import data files stored on a web server.
OVITO is now also available as an Anaconda package on Windows.
Improvements to the Adjust View dialog, which now allows interacting with the viewports while being open.
Bug fix: Installing a third-party extension module in ovitos fails on macOS due to security restrictions (issue #191).
Bug fix: Checkbox “File contains multiple timesteps” in GUI does not reflect correct state of file reader.
Integrated PM Larsen’s scheme of calculating the centrosymmetry parameter using minimum-weight matching.
New crop option in Viewport.render_image() Python function.
Windows installer is now digitally signed to be compatible with Microsoft Authenticode.
The embedded Python interpreter ovitos now ignores PYTHONPATH environment variable and user package directories (issue #189).
Fixed LAMMPS data file export for simulation cells aligned along negative coordinate axes (issue #188).
New algorithm type in Common Neighbor Analysis modifier: Interval CNA
Bug fix: Crash during animation frame change when using DXA modifier.
New “Denoising filter” option in OSPRay renderer, which greatly improves the visual quality of rendered images.
New “Every Nth frame” option in animation settings dialog.
Combined
bin_count_{xyz}parameter fields ofSpatialBinningModifierclass into singlebin_countfield.GSD file reader: Added support for the SphereUnion particle shape specification (see here).
Fix of UI bug: Data inspector panel not opening correctly in some situations.
OVITO now available as a Conda package for Linux and macOS.
Extended the Viewport.zoom_all() Python function.
Fixed broken (since dev679) OVITO program package for Windows, which crashed when inserting modifiers into the pipeline.
Bug fix: Pipeline stuck in infinite update loop when setting fractional playback rate for trajectory (issue #181)
Added Python function ovito.enable_logging(), which lets OVITO print activity information to the terminal during long-running operations.
New implementation of OVITO’s asynchronous task framework and pipeline execution/caching system.
New modifier: Time averaging
New modifier: Smooth trajectory
New option to load all frames of the trajectory into memory.
Modifiers, viewport layers, and pipelines can now be given user-defined names in the pipeline editor.
Data tables can now be created from Python, e.g. to have custom analysis modifiers that generate data plots.
Simplified usage of the Unwrap trajectories modifier, which now scans the input trajectory automatically in the background.
Changed default color scheme for unnamed particle types.
Updating the trajectory from the external file will automatically jump to the end of the trajectory.
Fixed ImportError in PyPI package on macOS (see this discussion).
Support for version 2.0 of the GSD file format (issue #176).
Fixed regression #175 in Expression Selection modifier, which was sporadically crashing since build dev476.
Now offering two separate program editions: OVITO Basic and OVITO Pro.
Added file reader for the oxDNA file format and a specialized visual element for nucleotides.
Removed the POV-Ray rendering engine (POV-Ray scene file export is still available).
Renamed DataSeries to DataTable throughout the scripting API, user interface and documentation.
Enhancements to the Cluster Analysis modifier: calculation of cluster centers of mass and unwrapping of particle coordinates.
Python interface: Introduced the Viewport::underlays stack. Redesigned the Viewport Layers command panel tab.
Voronoi Analysis modifier provides new option to visualize the computed Voronoi cells.
Workaround for video encoding issue resulting in invalid MP4/MOV files for frame rates 2/4/8/16 fps.
Updated Qt libraries to version 5.12.6.
The Color Coding and the Assign Color modifier can now operate on surface meshes.
Bug fix: Segfault in Combine Datasets modifier second first dataset contains bonds but first doesn’t (issue #173).
Bug fix: Segfault when accessing a mutable sub-object field (e.g.
Particles.bonds_) whose value is None (issue #172).Construct Surface Mesh modifier: New option to transfer particle properties to generated surface mesh.
Replaced video encoding component Libav with FFmpeg, bringing high-quality animated GIF rendering.
Fixed issue that prevented relative paths to external data files in a .ovito state file to get updated when files are moved.
Standalone Python package with the ovito module now available at https://pypi.org/project/ovito/.
Replaced PyQt5 with PySide2 Python module.
Fixed loading of multi-frame PDB files.
XYZ file exporter now includes cell origin in header line if non-zero.
New Use mesh color option for particle types, which renders particles having a user-defined shape with the original mesh colors.
Added keyboard shortcut for the New Program Window function.
Added the
SimulationCell.delta_vector()andParticles.delta_vector()methods for calculating the vector connecting two particles.Added the
Particles.create_particle()andBonds.create_bond()Python methods for adding new particles and bonds to a system.Let the LAMMPS data file parser accept additional spaces between header keywords.
Included missing Tcl/Tk support files of Python interpreter in the Windows installation package.
Bug Fix: Affine Transformation modifier does not transform dislocation lines.
New option in the Slice modifier to visualize the plane in rendered images.
Added a file writer for the GSD/HOOMD file format.
Added a visualization element for voxel grids computed by the Spatial Binning modifier or imported charge density fields.
Implemented the QuickSurf algorithm in the Construct Surface Mesh modifier as an alternative surface generation algorithm.
Added a file reader for the CIF format (Crystallographic Information File).
Extended sections on data manipulation and user-defined modifier functions in the Python documentation.
GSD file reader reads particle shape definitions (ellipsoids, polygons, convex polyhedra, general 3d meshes).
Create Bonds modifier remembers if the user explicitly re-enables the display of an unusually large number of bonds.
GSD file reader supports user-defined per-particle, per-bond and global data chunks.
Upgraded build environment to Qt release 5.12.5.
Settings for animation FPS and rendering resolution are retained across program session.
Added graphics export function for data series plots.
Display of bonds now takes into account particle radii when calculating the length of half-bond cylinders.
Documented the Clone pipeline function in the user manual.
Use Ctrl/Command key modifiers to zoom all viewports to scene extents at once.
Renamed the Correlation function modifier to Spatial correlation function modifier.
Replaced calls to FFTW library with more lightweight KISS FFT library.
CASTEP .md file reader: Automatic conversion from Bohr to Angstrom units.
Reimplemented how animation timeline is adjusted to accommodate loaded trajectories; redesign of the animation settings dialog.
Bug fix: Memory footprint continuously increases during animation rendering.
Fixed bug in CASTEP MD file reader not recognizing
<- hvlines.New axis style option in coordinate tripod viewport overlay.
Fix for ovitos.exe error “unable to find Qt5Core.dll on PATH” on Windows.
New file exporter for VTK voxel grid files. Allows to export results of Spatial Binning modifier.
Set up automatic mapping to particle properties for 4-column XYZ files.
New Constrain Rotation option in the viewport context menu to turn on/off camera alignment with z-axis.
Maximized state of active viewport is kept across program sessions.
Bug fix: Save as defaults function in particle type editor not working for numeric particle types (issue #157)
Bug fix: export_file() Python function always performs pipeline evaluation at frame 0.
Added the Viewport.create_widget() Python method, which allows embedding an OVITO viewport into a PyQt5 GUI.
Bug fix: LAMMPS data exporter not writing dipole atom style files correctly.
Extended LAMMPS data file reader to support a wider range of atom styles, including hybrid.
Extended the GSD file reader to parse the periodic image information, which can be used by the Unwrap Trajectories modifier to unfold particle positions.
Added a file reader for ParaDiS data files containing discrete dislocation lines.
The Unwrap Trajectories modifier can now undo the cell flipping performed by LAMMPS to keep the tilt factors within certain limits.
New modifier: Chill+ algorithm for identifying water structures.
Added support for user-defined particle shapes.
Added file reader for the OBJ and STL formats, which store triangle meshes.
Extended the Construct Surface Mesh modifier to identify disconnected regions of the filled volume.
Added the Viewport.camera_up parameter, which gives control over the orientation of the vertical axis in rendered images.
New implementation of the surface mesh data structure, now supporting assignment of arbitrary properties to the vertices, faces and spatial regions enclosed by surface mesh.
OSPRay rendering engine is now available in Windows builds of Ovito.
Added the
DataCollection.apply()Python method, which allows directly applying a built-in modifier to a dataset.Updated the Gaussian Cube file reader to auto-detect Bohr/Angstrom units and support files containing multiple field values.
Extended PDB file reader to support multi-frame trajectory files.
Added option to POSCAR file writer for outputting atomic positions and velocities in reduced cell coordinates.
Added new option to Construct Surface Mesh modifier that allows selecting all atoms on the constructed surface.
Extended the Python interface of the Color Coding modifier to simply definition of custom color gradients.
The Wigner-Seitz modifier now supports loading varying reference configurations, which depend on the current timestep.
Improved appearance of axis tripod viewport overlay when looking head-on to an axis (issue #88)
Added the new Unwrap trajectories modifier.
Bug fix: XYZ files with varying numbers of named atom types (issue #137)
Bug fix: Globbing algorithm did not recognize file sequences such as file-0.001, file-0.0001, etc. correctly.
Create Bonds modifier now defines a new bond type and assigns it to all newly created bonds
Reimplemented the Generate Trajectory Lines modifier, now supporting systems with varying number of particles. Unwrapped trajectory lines can now optionally be displayed in wrapped form.
Updated version of the PTM modifier, which now supports OVITO standard reference orientations for computing crystal orientations. See merge request.
Built-in SSH client now supports servers that use two-factor authentication and keyboard-interactive authentication.
Basic support for Quantum Espresso data file format.
Added file reader for GALAMOST file format.
Added an enabled property to viewport overlay Python class, which allows users to temporarily turn off individual overlays.
Changed the signature of user-defined modifier functions. Instead of separate input and output data collections, a user-defined modifier function now takes just a single data collection, which can be modified in-place by the function. A fallback to the old function signature is implemented to maintain backward compatibility.
Per-type masses are now read from LAMMPS data files.
Bug fix: Elastic Strain Calculation modifier yields off-diagonal elements of atomic Green-Lagrangian strain tensor with wrong sign.
Fixed UI issue #119.
Voronoi analysis modifier now dynamically adjusts the number of output columns of the Voronoi Index property to the maximum face order.
Renamed the “Bond-Angle Analysis” modifier to “Ackland-Jones Analysis” modifier.
Renamed the “Bin & Reduce” modifier to “Spatial Binning” modifier.
Extended the Spatial Binning modifier to support 3d grids.
Fixed bug in the marching cubes algorithm used by the Create Isosurface modifier, which sometimes let the surface construction fail with an error message.
Added file reader for the DL_POLY format.
Added calculation of partial (element-wise) RDFs to the Coordination Analysis modifier.
Generalized the Compute Property modifier, which can now compute bond properties too. Furthermore, neighbor terms can now reference properties of the central particle as well.
Added computation of height-difference correlation function to Correlation Function modifier.
Removed the Compute Bond Lengths modifier from the program. The extended Compute Property modifier provides a similar functionality.
Lifted 1 billion atom limit of the LAMMPS, CFG and IMD file parsers.
Fixed program crash at program startup due to unhandled exception in case the OpenGL initialization fails.
Generalized the Manual Selection modifier, which now supports selecting bonds too.
Bug fix: Move plane to simulation box center function of Slice modifier not calculating the distance value correctly for non-unit normal vectors
Implemented correct 2D shear and volumetric strain calculation in Atomic Strain modifier for two-dimensional system.
Added the Transparency bond property and partial support for rendering semi-transparent bonds.
Added a Color particles by type option to structure identification modifiers such as CNA and PTM, providing the option to preserve the current colors of particles.
Added a Sort particles by ID option to some file readers, automatically sorting the imported particles w.r.t. unique identifiers to obtain a stable ordering.
Bug fix: Correlation function modifier crashes when simulation cell is smaller than selected FFT grid spacing (issue #106).
Bug fix: Atomic Strain modifier ignores turning off the Select invalid particles user option.
File sequence globbing now works for hidden files (issue #98)
Modifiers no longer overwrite global attributes with the same name produced by upstream modifiers in the pipeline. Instead, new unique attribute names are generated whenever needed.
Renamed the ovito.DataSet Python class to Scene.
The VASP file reader can now handle more variants of XDATCAR files, e.g. those with varying cell size
Animation playback direction can be reversed using the Shift keyboard modifier
New SSH client for loading files from remote hosts, which supports public key authentication and a wider range of key exchange methods.
Bug fix: File descriptors do not get closed when importing new files due to bug in GSD file I/O layer.
The Expression Selection modifier now supports selection of other kinds of elements, for example bonds, in addition to particles.
The Polyhedral Template Matching (PTM) function has been extended and can now identify diamond structures.
The Generate Trajectory Lines utility has been replaced with the new Generate Trajectory Lines modifier.
Viewport overlays now provide a background option, putting them behind the three-dimensional scene.
OVITO now reads Gaussian Cube files (atoms and voxel grid data).
The new Data Inspector panel has been introduced. It replaces the old Inspect Particles and Inspect Dislocations utilities.
“Display objects” are now called “visual elements” within OVITO.
Replaced the
Viewport.render()Python method with the newViewport.render_image()andViewport.render_anim()methods. TheRenderSettingsand theAnimationSettingsclasses have been removed from the Python interface.The Combine Particle Sets modifier has become smarter: Particle types from the two input datasets are now merged and type IDs are automatically remapped.
The LAMMPS dump and XYZ file exporters now provide control over the output precision for floating-point numbers.
Added a new option to the Wigner-Seitz defect analysis modifier, which allows retaining the current atomic configuration instead of the reference configuration.
Implemented automatic detection of multi-timestep files (for LAMMPS dump and XYZ formats).
Added Python bindings for the CorrelationFunctionModifier.
Added a file reader for the XSF data format of XCrySDen (atomic structures as well as 3d voxel fields are supported by OVITO).
Added Python bindings for the CoordinateTripodOverlay.
Modifier templates can now be exported/imported to transfer them between computers.
Added the new OSPRay rendering engine.
Added handling of out-of-range atom serial numbers in PDB files with more than 99,999 atoms.
Bug fix: Stack overflow in Tachyon renderer when rendering a large number of semi-transparent objects
Bug fix: Made calls to the FFTW3 library from the Correlation Function modifier thread-safe
Reimplemented the displacement vector calculation in the Atomic Strain modifier to fix an error that occurred when the cutoff radius was larger than half the simulation cell size.
Added a file writer that can export dislocation lines to the VTK file format used by ParaView.
Extended the dislocation inspection utility to display a dislocation’s start and end vertex position.
Added support for 64-bit integer particle properties and large datasets with 2+ billion particles (only usable from analysis scripts, rendering won’t work for more than 2 billion particles).
Changed internal floating-point type from float to double, i.e., all calculations will now be performed with double precision. Memory footprint of the program increases because of this.
Performance improvement: The Wigner-Seitz defect analysis now makes use of all processor cores (issue #50).
Bug fix: Progress bar is now updated during file export.
Added a file writer for the binary NetCDF format according to the AMBER convention (issue #41).
Thanks to a complete redesign of the data pipeline system, modifiers can now access the entire input trajectory, making it possible to implement temporal analyses that operate not just on instantaneous simulation snapshots. A first, simple example is the new Interpolate trajectory modifier, which can interpolate particle positions between simulation frames to create smooth looking animations from coarse sequences of simulation snapshots.
Furthermore, modifiers such as Atomic strain, Displacement vectors and Wigner-Seitz defect analysis can now use frame 0 of the currently loaded simulation sequence as reference configuration. It is no longer necessary to load the reference configuration from a separate input file.
Several modifier have been generalized and can now operate on other forms of data in addition to particles, e.g. bonds and their properties. Examples are the Assign color, Invert selection, Select Type, Histogram, Scatter plot, Replicate and Delete selected modifiers.
The Python programming interface has been redesigned and extended. See this page for more information.
Surface meshes, dislocation networks and voxel data grids now carry their own periodic domain information with them. Changing the master simulation cell size no longer screws up the display of these data types.
The LAMMPS binary dump file reader automatically detects the endianess used in the file. It can now read files produced on IBM BlueGene/Q machines, for example (issue #23).
The Correlation function modifier now exports radial distribution function and structure factor as extra columns to a text file.
A VTK file writer has been added, which can export surface triangle meshes produced by the Construct surface mesh and Create Isosurface modifiers.
The ovitos script interpreter now accepts (and silently ignores) the -u command line option of CPython for better compatibility with pip install scripts that use this option (issue #43).
The Tachyon renderer now obeys the maximum number of parallel threads set by the
--nthreadscommand line option.The Show Periodic Images modifier has been renamed to Replicate modifier. In addition to particles and bonds it now can replicate surfaces, dislocation lines and voxel data grids too (issue #39).
Version 2.9.0 (27-Jul-2017)
Added the
--nthreadscommand line option to ovitos as an alternative to-nt(issue #35).Brought back missing stderr output from calls to sys.exit() in ovitos interpreter.
Extended the Particle Inspection utility to allow expression-based selection of particles in addition to picking them using the mouse (issue #19).
Added a bond-based mode to the Cluster Analysis modifier. It allows forming clusters based on the bond network topology.
The OVITO main window now accepts data files and .ovito files via drag & drop (issue #28).
Extended the Load Trajectory modifier to also copy other varying particle properties in addition to the particle positions (issue #29).
The particle indices displayed by the Particle Inspection utility are now zero-based, consistent with the ParticleIndex variable used by the Expression Selection modifier (issue #21).
Added the new Relative face area threshold parameter to the Voronoi analysis modifier, which allows filtering out small faces with an area below a specified fraction of the total Voronoi cell surface area (issue #7).
Added file parsers for the CASTEP .cell, .geom and .md simulation file formats.
Replaced the eliminate homogeneous deformation option of the Displacement Vectors modifier with the more general affine mapping setting.
On macOS, data files and .ovito files can now be associated with OVTIO and directly opened from the Finder (issue #22).
Added the FileSource.loaded_file attribute, which allows accessing the filename of the currently loaded simulation file from a Python script.
New modifier: The Voronoi Topology Analysis modifier can classify the Voronoi polyhedra of particles, e.g. to perform structural filtering.
New modifier: The Correlation Function modifier has been contributed by Lars Pastewka. It allows computing the spatial correlation between two particle properties.
Bug fix: LAMMPS data file parser ignored Bonds section at end of file when number of bonds is zero
New modifier: The Create Isosurface modifier allows visualizing field quantities like the electron density that are defined on a structured data grid. So far, only the POSCAR file parser has been extended to read charge density data from CHGCAR files, which can serve as input for the isosurface modifier.
New modifier: The Coordination Polyhedra modifier constructs convex hulls from the bonded neighbours of atoms.
Bug fix: Unexpected error message during file export when the old mapping for the output file columns has become invalid.
Added the Adjust range (all frames) function to the Color Coding modifier, which takes into account all frames of the animation sequence when determining the min/max values of the input property.
Added a new user option to the Affine Transformation modifier that enables the transformation of vectorial particle properties like Force and velocity together with the particle positions (issue #11).
Bug fix: Need to pass /exit option to Windows version of POV-Ray to automatically close message window after rendering is done (issue #16).
Bug fix: Assertion error in Ambient Occlusion modifier when modifier input is empty.
The ovito Python module is now usable from external Python interpreters as well, not only ovitos.
Complete overhaul of the internals of the asynchronous data pipeline framework; many improvements to the code.
Dropped backward compatibility with Qt library versions less than 5.4.
Bug fix: Parser error when reading a LAMMPS file containing very small numbers on the order of 1e-200 (issue #12).
Bug fix: Windows version always appends .pov to expected file names (issue #13).
Bug fix: Create trajectories function does not use correct number of frames from input sequence (issue #6).
Added support for omnidirectional stereoscopic rendering to the POV-Ray renderer plugin. This allows producing 360 degrees VR movies (requires POV-Ray 3.7.1).
The required system library libstdc++.so.6 is no longer bundled with the Linux version of OVITO, because it causes conflicts with OpenGL drivers on some systems.
Reading multi-frame GSD files that contain static data now works correctly.
Bug fix: Program crash during parallel access to NetCDF files. Calls to NetCDF library functions are now serialized, because they are not thread-safe.
Version 2.8.2 (24-Jan-2017)
The Histogram modifier can now compute the distribution of bond properties too (e.g. bond lengths).
Modifiers now report an out-of-memory condition. Ovito no longer crashes when a memory allocation fails during modifier evaluation.
Fixed viewport rendering and other issues for simulation datasets with a very small length scale (~10-11).
Added Python bindings for particle trajectory line generation and visualization.
Version 2.8.1 (17-Dec-2016)
Bug fix: Segmentation fault after applying a Color Coding modifier with Viridis color map to particle data containing NaN values.
Bug fix: Segmentation fault when closing Scatter Plot editor panel.
Bug fix: Compute Property modifier failed with an error when preceded by a Dislocation Analysis (DXA) modifier, due to expression variable names containing invalid characters.
Added read/write support for LAMMPS data files with atom_style sphere.
Added the NearestNeighborFinder.find_at() Python method for querying the nearest particle(s) around an arbitrary spatial position.
Added a detailed usage example to the scripting documentation of the WignerSeitzAnalysisModifier to demonstrate the identification of specific point defect types, e.g. antisites.
Bug fix: Viewport.render() Python method failed in GUI mode with an error.
Bug fix: FileSource.load() Python method failed when called with keyword arguments.
Bug fix: Serial computations performed by the data pipeline occupy two processor cores instead of one.
Bug fix: A custom modifier script function, raising an exception and running as part of a data pipeline within ovitos, produced assertion error.
Bug fix: Cyclic reference between DataSet and ScriptEngine classes led to a potential memory leak when using Python script modifiers or viewport overlays within a script executed by ovitos.
MacOS version of OVITO is now distributed as a signed application bundle in the form of a DMG disk image (avoids “unidentified developer” warning message on first start).
Disabled geometry shaders by default for AMD/ATI hardware on Windows due to compatibility problems reported by some users.
Python modifier and Python viewport overlay scripts can now be edited in a separate code editor window.
Version 2.8.0 (23-Nov-2016)
Added the POV-Ray rendering backend and the POV-Ray scene file exporter.
Fully transparent, invisible particles are no longer sent to the Tachyon renderer to avoid artifacts.
Replaced QCustomPlot component with QwtPlot component for graph plotting within the GUI.
Replaced Boost.Python with pybind11 library to implement OVITO’s Python bindings.
The DXA modifier now outputs attributes for the computed line lengths which are broken down by dislocation type.
Bug fix: Segmentation fault in Color Coding modifier when input particle property contains infinite values.
Fixed an issue with the Freeze Property modifier, which didn`t attach a display object to vector properties.
The “Fusion” UI theme is now explicitly activated on Linux. Older builds of OVITO used the “Windows” theme for some reason.
Bug fix: Atomic strain calculation failed for 2d simulation cells with zero length along Z.
The Linux binaries are now built with the gcc 5.1 compiler.
Bug fix: Identify diamond structure modifier did not output the structure counts as global attributes.
Removed restriction of LAMMPS binary dump file reader to less than 200k atoms.
Added a command line option to the ovitos program, which gives users control over the number of parallel threads used by OVITO.
Added depth-of-field rendering to the Tachyon renderer.
OVITO can now color dislocation lines based on their Burgers vectors (only BCC crystals so far). Before, coloring was possible only on the basis of dislocation type or dislocation character.
Added the Generate perfect dislocations option to the DXA modifier, which suppresses the identification of partial dislocations.
Improved I/O performance of the LAMMPS data file reader when reading bonds information.
Made PDB file reader compatible with files generated by the Gromacs trjconv tool.
Extended the Color Coding modifier to support the coloring of vector arrows (in addition to particles and bonds).
Bug fix: If the Compute Property modifier is used to create a vector property, the display settings for vector arrows are no longer lost every time the modifier is re-evaluated.
The Text Label and Color Legend viewport overlays can now draw an outline around text to make it easier to read on all backgrounds.
Disabled some strict conformance checks in the SSH module to improve compatibility with some SSH servers.
Bug fix:
Viewport.render()Python function raised a Boost.Python.ArgumentError when being called in GUI mode.Bug fix: NetCDF file parser did not recognize unwrapped particle coordinates in files written by LAMMPS.
Raised the threshold values for the automatic selection of particle rendering quality. High quality mode is now being used for <4,000 particles, medium mode for <400,000 particles, and low quality mode for everything above.
OVITO now ships with the matplotlib Python module. This makes it possible to integrate custom data plots into images or movies rendered by OVITO.
Bug fix: Unicode characters are now correctly rendered by the Color Legend viewport overlay.
Added Python interface for the FreezePropertyModifier.
Added two additional colormaps to the Color Coding modifier: Magma and Viridis.
Added a Combine particle sets modifier, which allows merging two datasets into one.
Added GSD (General Simulation Data) file reader for HOOMD-blue simulation files.
Bug fix: Inserting a modifier while scanning an XYZ/LAMMPS input file to discover simulation frames may crash program.
The ovitos interactive interpreter now creates/loads a separate IPython profile named ovito instead of the default profile. This reduces inteference with an existing IPython installation on the same system.
Workaround: Exporting denormalized floating-point numbers to an output file crashes program due to bug in Boost.Karma library.
Regression: Compatibility with high-resolution displays (Retina/Mac OS)
Regression: OpenGL rendering does not work in console mode on Windows
NetCDF reader now accepts files where particle data is stored in a subgroup named AMBER.
Version 2.7.1 (28-Aug-2016)
Integrated IPython in Linux and Mac OS builds of OVITO.
Small bug fix in Animation Settings dialog: Changing the frame rate made the time slider jump.
Updated the NetCDF file reader to make it compatible with files written by the SimPARTIX code.
Updated the VTK file reader to accept empty lines in the header.
Updated the IMD file reader, which did not correctly map file columns to standard particle properties with vector components.
Updated the CFG file reader to accept empty lines preceding the first header line.
Updated the PDB file reader to parse molecule identifiers and types.
Added the
ParticleTypeProperty.get_type_by_id()andget_type_by_name()methods to the Python interface.Added the DislocationSegment.spatial_burgers_vector property to the Python interface.
Added a confirmation message before resetting the URL history list in the SSH connection dialog.
Version 2.7.0 (25-Jul-2016)
Bug fix: Program crashed when entering a non-valid text into a numeric input field.
Added OpenGL driver-bug workaround to fix high-quality rendering of particles on Linux/Intel graphics systems.
Improved visual appearance of large number of semi-transparent particles in images generated by the Tachyon renderer.
Fixed a bug in the line coarsening routine of the DXA modifier.
Periodic image shift vectors of bonds are now accessible from Python scripts.
The Atomic Strain calculation modifier can now perform a polar decomposition F=RU of the deformation gradient into a rotation and stretch tensor.
Added a Use only selected particles option to the Histogram and Bin and Reduce modifiers, which lets you to restrict the calculation to a subset of particles.
Extended the Color Coding modifier so that it can also operate on bonds instead of particles.
Added the Compute Bond Lengths modifier, which can be used in conjunction with the Color Coding modifier to color bonds according to their length.
Added the Text Label viewport overlay, which provides an easy way of inserting a text label into rendered images and movies. The text can contain placeholders which are replaced with quantities computed by OVITO.
Added a text file export function, which allows exporting scalar quantities computed by OVITO to a tabular text file as functions of simulation time.
OVITO can now load XDATCAR trajectory files written by Vasp.
Added the Polyhedral Template Matching modifier, which can robustly identify lattice structures at high temperature.
The LAMMPS dump file exporter now writes out the simulation timestep number read from an input dump file to the file header instead of the animation frame.
Added the Load Trajectory modifier, which allows loading datasets that consist of separate topology and trajectory files (e.g. a LAMMPS data file with bond definitions and a LAMMPS dump file with atom trajectories).
The pair-wise cutoff mode of the Create Bonds modifier is now usable from Python.
Improved visual quality of particle display for very distant and small (sub-pixel) particles.
Added a lower cutoff parameter to the Create Bonds modifier.
Scene files that refer to external data file in the same directory can now be transferred to another computer without breaking the link.
Documented the CA dislocation file format in the user manual. This makes it possible to export the extracted dislocation lines and process/analyze them outside of OVITO.
Fixed a bug in the Create Bonds modifier, which produced wrong results when all pair-wise cutoffs were set to values smaller than 1.0.
OVITO no longer depends on the CGAL library, making it easier to build it from source.
The CA dislocation file importer now supports multi-timestep files.
Fixed a bug in the Show Periodic Images modifier, which did not replicate bond properties.
The behavior of the
ovito.io.import_file()Python function has been changed. It no longer adds the created node to the scene, and it is now okay to call this function repeatedly. It also accepts wildcard filenames now to import file sequences. See the updated Python documentation for details.The
ovito.io.export_file()Python function now gives full control over which animation frames are being exported.Fixed bug in the Coordination Analysis modifier, which prevented the RDF calculation for 2D systems.
Version 2.6.2 (19-Mar-2016)
Updated the built-in SSH client to support more encryption methods and improve compatibility with some SSH servers.
Small bug fix that solves a (rare) problem with the display of dislocation lines in periodic systems.
For very small, periodic simulation cells the Elastic Strain Calculation modifier silently failed for diamond crystals. Now the user is informed that the Show Periodic Images modifier should be applied first to extend the box size.
Fixed output of animated GIF files on Linux.
Reduced memory footprint of the DXA analysis modifier.
The Slice modifier now also cuts surface meshes generated by the Construct Surface Mesh modifier and dislocation lines generated by the DXA analysis modifier.
Introduced basic support for 2D systems. The 2D flag can be set in the simulation cell properties panel. The Atomic Strain and Coordination Analysis modifiers then perform the computation in 2D (i.e. the XY plane).
Key-value pairs read from extended XYZ file headers and the LAMMPS timestep number are now accessible from Python via the new DataCollection.attributes property.
Added the Number of bins parameter to the Coordination Analysis modifier, allowing the user to control the resolution of the generated RDF histogram.
Errors generated by Python scripts that are run via the Run script file function are now displayed in the GUI.
Re-added the Every nth frame field to the Render Settings panel, which allows rendering only a subset of animation frames.
Fixed import of Aspherical Shape particle property values from a file.
The export_file() script function can now produce multi-timestep files.
Improved automatic detection of PDB files and added parsing of bonds defined by CONECT records.
Fixed bug in Elastic Strain Calculation modifier, which got stuck in infinite loop when applied to some complex polycrystalline structures.
Moving the viewport camera no longer stops animation playback.
Added the Vector Color particle property, which allows changing the display color of vector arrows on a per-particle basis.
Bug fix: The Create Bonds modifier reported the number of half-bonds, not the number of full bonds generated.
Regression: Restricting the Compute Property modifier to selected particles did not work correctly; existing values for unselected particles were always reset to zero.
Added the CA file exporter, which allows saving DXA analysis results to disk (and load them again at a later time).
Version 2.6.1 (15-Nov-2015)
Arrows can now be centered on particles, e.g. to visualize magnetic moments and other vector properties. Atomic force vectors and dipole vectors read from simulation files can now be directly visualized.
The Tachyon renderer now supports transparent backgrounds.
The Color Legend and Coordinate Tripod viewport overlays can now be repositioned with the mouse.
Display objects that are generated by modifiers no longer temporarily disappear from the pipeline editor list while playing an animation.
The PDB file parser now accepts lines with up to 83 characters to support files written by Accelrys Discovery Studio.
Fixed regression in CA file importer, which is broken in previous release.
Version 2.6.0 (02-Nov-2015)
Added the <a+c> dislocation type for HCP crystals to the Dislocation Analysis modifier.
Added the Elastic Strain Calculation modifier, which computes the atomic-level elastic strain and deformation gradient tensors in crystalline systems. It can be used to analyze local elastic distortions in a crystal lattice and to determine the local crystal orientation.
The modification pipeline editor now allows changing the application order of modifiers via drag and drop.
Added the Dislocation Inspection utility, which lets you obtain further information about dislocation segments extracted by the Dislocation Analysis modifier.
Added an introduction to OVITO’s animation system to the manual, which explains how to animate parameters and the camera.
The Compute Property modifier can now perform computations that take into account the local neighborhood of particles. For example, you can use this to average a quantity over spherical regions around each particle.
The Wigner-Seitz analysis modifier can now break down the computed site occupancy number into per-type numbers. This makes it easier to identify antisite defects, for example.
Viewports are no longer automatically zoomed to show everything when replacing the currently loaded dataset with a new one. This makes it easier to preserve the current view configuration when switching to a different simulation file.
Fixed the assignment of dislocation lines that were loaded from a CrystalAnalysis file to Burgers vectors families. The CA file import was broken since version 2.4.4.
Added a bond-based mode to the Common Neighbor Analysis modifier, which computes the CNA indices based on existing bonds between particles. The modifier also outputs the computed CNA bond indices as a new bond property. This enables, for example, analyses of disordered systems using the classical CNA.
It is now possible to write you own modifiers in Python.
Added a Use only selected particles option to the Common Neighbor Analysis and the Identify Diamond Structure modifiers, which allows identifying sub-lattices.
Added the NearestNeighborFinder Python class, which can be used from Python scripts to find the N nearest neighbors of a particle.
Python scripts can now access the dislocation lines extracted by the DislocationAnalysisModifier.
Bug fix: Bond properties like the bond type are now updated when dangling bonds are removed due to deleted particles.
Fixed bug in Atomic Strain modifier, which produced weird results when PBC flags of reference configuration do not match PBC flags of deformed configuration. Now PBC flags of deformed configuration always override the boundary conditions of the reference simulation cell.
Fixed error in ovito.dataset.scene_nodes.__iter__() Python method.
Fixed bug in file parser that led to wrong particle type names when loading a multi-frame XYZ file with varying set of named atom types.
Bug fixes in Python-ASE interface – Cell matrix is transposed and duplicate properties are handled.
Version 2.5.1 (07-Aug-2015)
The LAMMPS data file exporter can now produce files with LAMMPS atom styles other than atomic. It also exports bonds if present.
Arbitrary triclinic simulation cells can now be exported to the LAMMPS data file format. They will be automatically transformed to the canonical LAMMPS representation.
The LAMMPS data file parser now reads bond types.
Added the fmod(A,B) math function to the Compute Property and Expression Select modifiers.
Added visualization support for cylindrical and spherocylindrical particles.
Added a file parser for FHI-aims log files, which can contain multiple simulation frames.
Added the Indicate line direction option to the dislocation display object.
Version 2.5.0 (25-Jul-2015)
Added Python interface for Bin and Reduce modifier.
Fixed viewport font issue on Macs with high-dpi display.
Added the Expand Selection modifier.
Added a No bonds between different molecules option to the Create Bonds modifier.
Changed the behavior of the IMD file exporter: All particle properties to be exported, including the standard ones defined by the IMD format, must now be explicitly selected by the user.
Integrated DXA (Dislocation Extraction Algorithm) into OVITO.
Voronoi analysis modifier can now output bonds between neighboring atoms which share a Voronoi face.
Python scripting interface now allows conversion to/from ASE Atoms objects.
Python scripting interface now supports write access to particle properties and procedural generation of input particle datasets for OVITO’s modification pipeline.
Bug fix: Large number of simultaneous SFTP download requests led to error message SFTP error: Server could not start session.
Bug fix: Using the Python viewport overlay led to program crash on Windows 8 x64.
Scripting function Viewport.render() now returns the rendered image, which can be manipulated before saving it to disk.
Certain modifier and display parameters can now be animated using animation keys.
Fixed OpenGL rendering of bonds/arrows on certain Windows/NVidia systems.
Added import/export support for FHI-aims file format.
Colors and radii used for particle types (as well as structure types) can now be predefined by the user.
Migrated to version 5.4 of the Qt library. This may affect OpenGL rendering and the viewport display. Please report any issues that you experience.
Bin & Reduce modifier now takes into account the simulation box origin, uses double precision numbers to perform calculations, and 1D-plot now spans the entire interval.
Bug fix: NetCDF file importer doesn’t close file handle, leading to error after loading several thousand frames.
Bug fix: LAMMPS data file parser stumbles over AngleTorsion Coeffs file section.
Visualization of particle trajectory lines.
Frequently used modifiers or combinations of modifiers can be saved (including modifier settings) for quick access.
Rendering of ellipsoidal and box particles with orientation.
Positioning the mouse over a bond shows its properties in the status bar.
Fixed initialization of the Select Particle Type modifier.
Fixed error when loading a compressed simulation file >2GB.
Version 2.4.4 (29-Mar-2015)
Fixed error when rendering a high-resolution video.
Surface mesh computed by ConstructSurfaceModifier can now be exported to a VTK file from Python.
Added Python class ovito.data.CutoffNeighborFinder, which enables access to particle neighbor lists from Python.
Particles and bonds are now rendered in chunks in the OpenGL viewports to work around a memory limit on some graphics hardware.
Bond cylinders are now rendered using a geometry shader if supported by the graphics card.
The IMD file exporter now lets the user select the particle properties to export (instead of exporting all).
The VTK triangle mesh importer now reads per-face color information.
Version 2.4.3 (02-Mar-2015)
Upgraded integrated script interpreter from Python 2.7 to Python 3.4. Please update your scripts to make them compatible with Python 3.
Added rendering support for particles with non-cubic, axis-aligned box shape (via Aspherical shape particle property)
Added a dialog box to the Affine Transformation modifier, which lets the user enter a rotation axis, angle, and center.
Removed cutoff option from Voronoi Analysis modifier in favour of a faster algorithm for orthogonal simulation cells, which is based on Voro++ container classes.
The Voronoi Analysis modifier now determines the maximum number of edges per face in the Voronoi tessellation and warns if it exceeds the truncation length of computed Voronoi index vectors.
OVITO can now load bonds from LAMMPS data files.
The Freeze Property modifier now works when particles are lost during the simulation.
Similarly, the Atomic Strain analysis can now deal with simulations where the number of particles is not constant.
The Wrap at Periodic Boundaries modifier now wraps bonds crossing a periodic boundary.
Added a scriptable viewport overlay, which allows painting custom text and graphics over the rendered image. See how you can use it to add a scale bar to a viewport.
The Show Periodic Images modifier now replicates bonds too.
The XYZ file import now displays the file’s comment line in the status field.
Switched from MinGW to Visual C++ 2013 compiler to build Windows version. Python scripting is now supported by the 64-bit program version for Windows too.
Removed old Javascript plugin.
Bug fix:
--versioncommand line option causes program to crash.Bug fix: XYZ file column mapping dialog showed the column names from the last loaded extended XYZ file.
Version 2.4.2 (14-Nov-2014)
The Color Coding modifier now supports user-defined color maps.
Significantly improved performance of cutoff-based neighbor finding and k-nearest neighbor search routines. This code optimization speeds up many analysis algorithms in OVITO, in particular for large datasets.
Added the Identify Diamond Structure analysis function, which finds atoms that form a cubic or hexagonal diamond lattice.
Dialog box asking to save changes is only shown when scene has already been saved before.
The Color Legend overlay now provides an option to overwrite the numeric labels with a custom text.
Bug fix: Periodic boundary flags were not correctly updated when loading a new file using the Pick new local input file button.
Bug fix: Viewport.render() Python function raised error when called without a RenderSettings object.
Version 2.4.1 (01-Nov-2014)
New integrated Python engine, which provides a powerful scripting interface (see scripting documentation). This is going to replace the Javascript engine, which has been deprecated and will be removed in a future program version. Command line options to run old scripts have been renamed to –jsscript and –jsexec.
New Voronoi analysis modifier, which can compute atomic volumes, coordination numbers and Voronoi indices.
It’s now possible to include the coordinate system tripod and a color legend in the rendered image.
Particle properties are displayed in the status bar when hovering over particles in the viewports.
Periodic boundary conditions can be overridden by the user without the changes being lost when a new simulation frame is loaded.
Added import/export support for extended XYZ format (see http://jrkermode.co.uk/quippy/io.html#extendedxyz), which includes metadata describing the data columns and the simulation cell.
Improved input and output performance for text-based file formats.
The OpenGL renderer can now display semi-transparent particles and surfaces.
Added calculation of non-affine displacements to Atomic strain modifier. (This is Falk & Langer’s D2min measure, see the 1998 PRB.)
New Bin and reduce analysis modifier.
The Create bonds modifier can now handle particles that are located outside a (periodic) simulation box.
The Color coding modifier can display a color legend in the rendered image.
Added a file parser for PDB files (still experimental).
Added basic keyframe animation support. Some modifier parameters and other settings can now be animated. Future versions will also offer camera animation capabilities.
The Show periodic images modifier can now assign unique IDs to particle copies.
LAMMPS data file parser now supports additional LAMMPS atom styles such as charge and bond.
Fixed high-quality particle rendering on Windows computers with Intel HD 4000 graphics.
Bug fix: Export of compressed LAMMPS data files could result in truncated files.
Bug fix: Solid volume computed by Construct surface mesh modifier could be inaccurate due to low numerical precision
Bug fix: Construct surface mesh modifier crashed with certain input data.
Bug fix: VTK mesh file parser couldn’t handle multiple points per line (as written by ParaView).
Bug fix: LAMMPS data file parser did not parse atom IDs.
Bug fix: Particle inspection utility did not recalculate displayed distances and angles upon simulation frame change.
Bug fix: StrainTensor.XZ and StrainTensor.YZ components output by Atomic Strain modifier were swapped.
Bug fix: Fixed issue in Histogram modifier that occurred when the x-range was fixed to an interval smaller than the value range.
Bug fix: Atom type ordering is now maintained when importing a sequence of LAMMPS dump files with named atom types.
Version 2.3.3 (22-May-2014)
Added user options to application settings dialog that provide control over certain OpenGL-related settings. This allows working around compatibility problems on some systems.
User can now choose between a dark and a light viewport color scheme.
Added scripting interface for Tachyon renderer.
Added support for NetCDF files with variable particle numbers and with named particle types.
Added user options that control the automatic fetching of the news page from the web server and the transmission of the installation ID.
Fixed bug in camera orbit mode, which was not correctly restricting the camera rotation for some coordinate system orientations.
Version 2.3.2 (07-Apr-2014)
Fixed bug in Wigner-Seitz analysis modifier, which could cause a program crash when numbers of atoms in reference and current configuration differ.
Version 2.3.1 (01-Apr-2014)
Added saving and loading of presets for file-column-to-property mappings.
Added the
--execcommand line option, which allows directly executing a script command or to pass parameters to a script file.When opening an XYZ file, the column mapping dialog now displays an excerpt from the file’s header to help the user figure out the mapping.
The Construct Surface Modifier no longer creates cap polygons if the periodic simulation cell contains no particles.
Version 2.3.0 (29-Mar-2014)
Added the new scripting interface, which allows automating tasks.
Added the Freeze property modifier, which can prevent a particle property from changing over time.
Added the Scatter plot modifier, which plots one particle property against another. This modifier has been contributed by Lars Pastewka.
Added the Wigner-Seitz analysis modifier, which can identify vacancies and interstitials in a lattice.
Added a file importer for NetCDF files. Code was contributed by Lars Pastewka.
Added more input variables to the Compute property and Expression select modifiers (e.g. reduced particle coordinates and simulation cell size).
It’s now possible to load a sequence of files with each file containing multiple frames.
Fixed bug in CFG file importer, which did not read triclinic simulation cells correctly.
Fixed shader compilation error on OpenGL 2.0 systems and some other OpenGL related issues.
Version 2.2.4 (29-Jan-2014)
Fixed particle picking issue on computers with Intel graphics.
Fixed OpenGL display issues on systems with Intel graphics.
Fixed blurred viewport captions.
Fixed program crash when changing particle radius/color without having selected a particle type first.
Version 2.2.3 (16-Jan-2014)
Fixed the CFG file importer, which can now read CFG files written by newer versions of LAMMPS correctly. Auxiliary file columns are automatically mapped to OVITO’s standard particle properties if possible.
Modified particle file importers to ensure stable ordering of particle types (using lexicographical ordering when atom types have names, and ID-based ordering otherwise). The ordering of named particle types is now independent of their first occurrence in the input file.
Improved compatibility with some OpenGL implementations (Intel HD graphics on Windows and ATI Mobility Radeon HD 5470).
A 64-bit version of the program is now available for Windows.
A construction grid can be displayed in the viewports.
Version 2.2.2 (05-Jan-2014)
Fixed the following regression: Rendering a video with OVITO 2.2.1 resulted in an empty movie file.
Fixed display of the polygon path when using Fence selection (Manual Selection modifier).
Version 2.2.1 (26-Dec-2013)
Added a file parser for binary LAMMPS dump files.
Added a dialog window that displays information about the system’s OpenGL graphics driver. This dialog can be accessed via the Help menu.
Fixed bug in the Expression Select and Compute Property modifiers, which couldn’t handle particle property names that start with a number.
The OpenGL compatibility profile is now used instead of the core profile on Windows and Linux platforms.
Fixed an issue in the Construct Surface Mesh modifier, which sometimes led to a program crash on Windows.
Version 2.2.0 (15-Dec-2013)
The Construct Surface Mesh modifier has been added, which builds a polygonal mesh around a particle set.
The Cluster Analysis modifier has been added, which decomposes a particle system into clusters.
A new experimental visualization module has been added, which allows working with data generated by the Crystal Analysis Tool and the Dislocation Extraction Algorithm (DXA).
The Coordination Analysis modifier can now export the computed radial distribution function to a text file.
Added a new user option to the application settings dialog that allows turning off the restriction of the vertical camera rotation.
Added the File->New Window menu item, which opens another OVITO window. This makes life easier on the Mac OS platform, where starting multiple instances of the application is difficult.
The XYZ file exporter now writes particle type names instead of numeric type identifiers.
Added help buttons to parameter panels, which open the corresponding page in the user manual.
The manual is now included in every installation package. An internet connection is no longer necessary to access the manual.
Fixed the rendering of particle markers.
Version 2.1.0 (15-Nov-2013)
The Manual Selection modifier has been added, which allows selecting individual particles with the mouse in the viewports. With the “Fence selection” mode, a group of particles can be easily selected by drawing a closed path around it.
OVITO is now able to display particles with cubic and square shape. This can be useful in visualizing large 2d lattice systems or Ising models.
OVITO now gives the user the option to import more than one dataset into the same scene and display them side by side.
A newly added VTK file importer allows reading triangle meshes to visualize geometric objects such as an indentor tip.
Camera objects can be created through the viewport context menu. A viewport can be linked to a camera object to show the corresponding view.
The OpenGL rendering code has been updated to better support older graphics cards and to improve compatibility with more graphics drivers.
The Tachyon renderer now supports semi-transparent particles. The transparency is controlled through the “Transparency” particle property. Use, for instance, the Computer Property modifier to set this property for certain particles. Transparency values can range from 0 (=fully opaque) to 1 (=not visible). Note that the interactive OpenGL renderer does not support transparency yet. Thus, all particles will still appear fully opaque here.
When importing a sequence of simulation snapshots, one can now configure the mapping of input frames to OVITO’s internal animation frames. This allows generating output movies with fewer (or more) frames than the imported snapshot sequence. This feature is in preparation for a future camera animation system.
The Mac OS version is now built against version 5.2 (beta) of the Qt library. This should fix a nasty UI bug on this platform (due to the old version of that library), which made text fields lose input focus.
Fixed saving/loading of the gradient type selected in the Color Coding modifier.
Fixed a program deadlock when dragging the time slider with the mouse after loading a file sequence from a remote location.
Version 2.0.3 (22-Oct-2013)
Ported Tachyon raytracing renderer from old OVITO 1.1.0 release. This software-based rendering engine allows producing images with high-quality shading and ambient occlusion lighting.
The Create Bonds modifier will automatically turn off the display of bonds when (accidentally?) creating a large number of bonds (1+ million), which would make the program freeze for at least several seconds.
The Displacement Vectors modifier now supports relative reference frames, i.e., displacements can be calculated from two snapshots separated by a fixed time interval. Before this addition, the modifier could only compute displacements with respect to a fixed reference simulation snapshot.
The Inspect Particle applet now lets one select multiple particles and can report distances and angles between particles.
Added Clear history button to remote file import dialog.
The POSCAR file exporter now writes the new file format, which includes atom type names.
Added support for computers with high-resolution (Retina) displays.
Fixed bug in the Affine Transformation modifier leading to recursive updates.
Version 2.0.2 (30-Sep-2013)
Fixed loading of multi-timestep files with names containing a digit.
Fixed import of CFG file with atom type information.
Version 2.0.1 (27-Sep-2013)
Fixed loading of file sequences based on wildcard pattern on Windows platform.
Replaced const arrays in GLSL shaders with uniform variables to support older Intel graphics chips.
Version 2.0.0 (25-Sep-2013)
Many changes, almost a complete rewrite of OVITO’s code base.