Create bonds
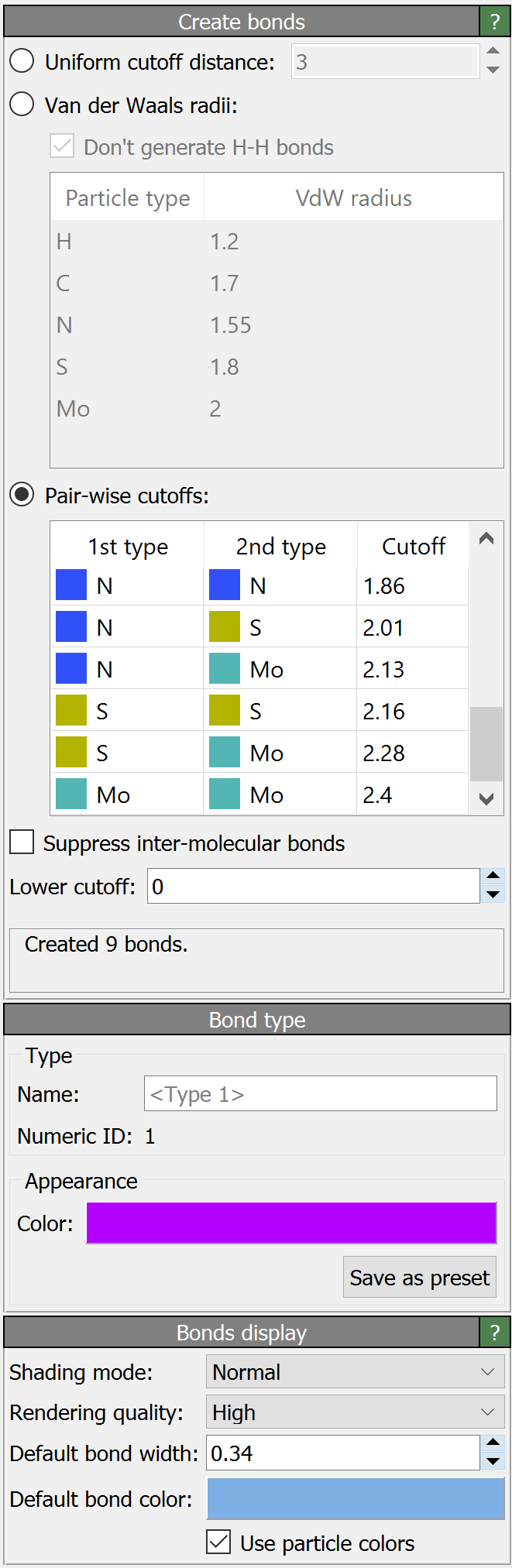
This modifier creates new bonds between pairs of particles according to a distance-based criterion. Existing bonds are not affected by the modifier and the modifier won’t create another bond between a pair of particles already connected by an existing bond.
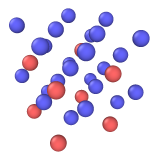
Input
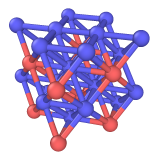
Output
Modes of operation
The modifier supports four modes of operation to determine which pairs of particles should be connected by bonds:
- Uniform cutoff distance
A uniform distance threshold is used to create bonds between pairs of particles. The species of the particles is not taken into account in this mode.
- Covalent radii
Bond creation follows a two-step process: First, an initial set of bonds is created based on a type-dependent distance criterion, for all pairs of particles that have a distance \(r_{ij} < r_{ij}^\text{Eq} + \epsilon\). Here, \(r_{ij}^\text{Eq}\) is the sum of the covalent radii of particles \(i\) and \(j\). These radii are taken from [J. Comput. Chem. 12, 891-898, 1991]. The padding \(\epsilon\) has a constant value of \(0.4\) (assuming Angstrom units).
This typically leads to some atoms with over-coordinated bonds. Therefore, a second processing step removes excess bonds again. Here, the maximum coordination tabulated in [J. Comput. Chem. 37, 1191-1205, 2016] is used as a target such that no atoms have excess bonds. To this end, the most elongated bonds relative to their equilibrium length are selected for removal.
To look up covalent radii and maximum coordination values, this method requires chemical particle types to be defined, i.e., the names of all particle types must match chemical symbols. Otherwise, the modifier will report an error and won’t create any bonds.
This bond creation algorithm is described in greater detail in the reference paper:
- Van der Waals radii
A bond is created between two atoms if their separation is less than 60% of the sum of their van der Waals radii. This criterion has been adopted from the popular visualization software VMD. The van der Waals radii of all particle types of the system are displayed in the table. OVITO initializes these values during file import based on the chemical element symbols found in the input file. If needed, you can override the standard van der Waals radius of each atom type using the Edit types modifier or, permanently, in the application settings. Bonds are only created between pairs of particles which both have a positive van der Waals radius.
Furthermore, the option Don’t generate H-H bonds is turned on by default, which means the modifier will not generate any bonds connecting two hydrogen atoms, i.e., atoms that both have a particle type named “H” - even if they fulfill the distance-based criterion.
- Pair-wise cutoffs
This mode gives you full control over the bond distance cutoff for each pair-wise combination of particle types. The table lists all pair-wise type combinations defined for the current system, and some of the cutoff values in the third column may already be pre-initialized according to the Van der Waals criterion described above. A positive cutoff value is needed to create bonds between pairs of particles of the given type(s). Note that this mode is only available if particle types have been defined for the system, i.e., the particle property
Particle Typeexists.
The option Suppress inter-molecular bonds restricts generation of bonds to particles that
are part of the same molecule, i.e., which have matching values of the Molecule Identifier property.
If the Molecule Identifier particle property is not defined for the system, this option has no effect.
The modifier lets you specify an optional Lower cutoff value. It effectively restricts the generation of bonds to the distance interval between the lower and upper cutoffs.
The modifier defines a new bond type, which is assigned to the newly created bonds. The properties of this bond type, in particular its name and display color, can be edited in the second parameter panel. Furthermore, the modifier will automatically create a bonds visual element if needed, which lets you control the visual appearance of bonds. The display settings of this visual element are found in the third parameter panel.
As for particles, OVITO supports the assignment of arbitrary properties to bonds which have been generated by this modifier or which were created during simulation file import. Certain bond properties (see this section) are used by OVITO to control the visualization of bonds. By changing the values of these properties, for example using the Compute property modifier, you can adjust the visual representation of individual bonds.
Attention
In case the modifier generated more than 2,000,000 bonds, it will show a warning message and automatically turn off the display of the bonds as a precaution, because rendering too many bonds in the interactive viewports of OVITO on a slow machine could take an exceedingly long time and freeze the entire program. If desired, you can manually show the bonds again by re-enabling the bonds visual element in the pipeline editor.
Technical notes
To correctly deal with periodic simulations, OVITO maintains a triplet of integer numbers with every bond, which is stored in the Periodic Image bond property array.
Each bond’s triplet specifies whether that bond crosses the periodic boundaries of the simulation cell (and in which direction).
For example, a bond crossing the periodic cell boundary in the positive X direction is indicated by the triplet (1,0,0) and
will be visualized as two separate half-bonds, one on either side of the simulation cell. Bonds in the interior of the simulation cell, which do not cross a
periodic boundary, have a Periodic Image value of (0,0,0).
See also
ovito.modifiers.CreateBondsModifier (Python API)